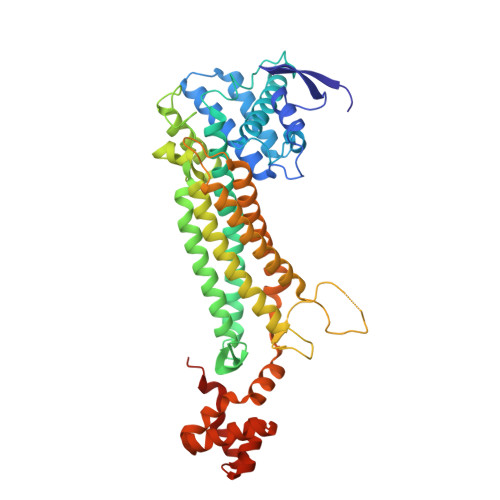
Closed fumarase C active-site structures reveal SS Loop residue contribution in catalysis.
Stuttgen, G.M., Grosskopf, J.D., Berger, C.R., May, J.F., Bhattacharyya, B., Weaver, T.M.(2020) FEBS Lett 594: 337-357
- PubMed: 31514245 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13603
- Primary Citation Related Structures:
6NZ9, 6NZA, 6NZB, 6NZC - PubMed Abstract:
Fumarase C (FumC) catalyzes the reversible conversion of fumarate to S-malate. Previous structural investigations within the superfamily have reported a dynamic structural segment, termed the SS Loop. To date, active-site asymmetry has raised the question of how SS Loop placement affects participation of key residues during the reaction. Herein, we report structural and kinetic analyses from Escherichia coli FumC variants to understand the contribution of SS Loop residues S318, K324, and N326. High-resolution X-ray crystallographic results reveal three distinct FumC active-site conformations; disordered-open, ordered-open, and the newly discovered ordered-closed. Surprisingly, each SS Loop variant has unaffected Michaelis constants coupled to reductions in turnover number. Based upon our structural and functional analyses, we propose structural and catalytic roles for each of the aforementioned residues.
- Department of Chemistry and Biochemistry, University Wisconsin - La Crosse, WI, USA.
Organizational Affiliation: