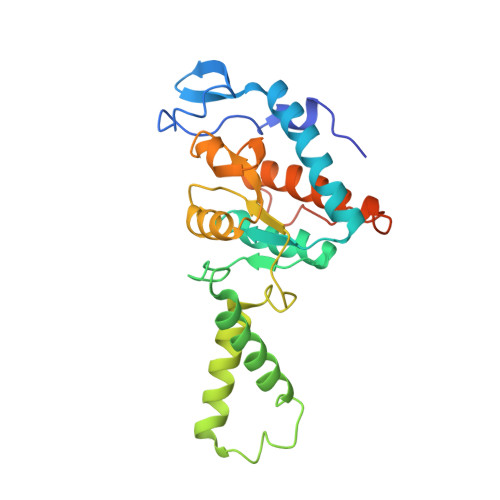
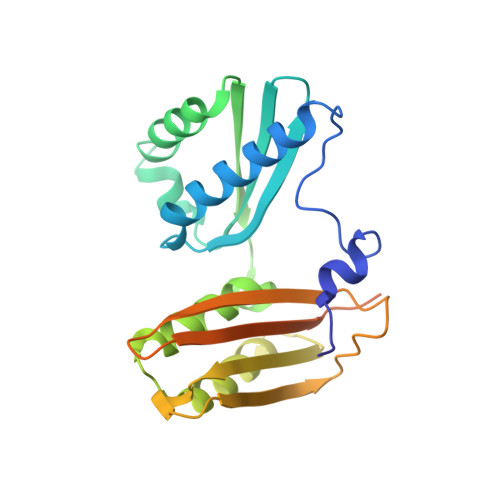
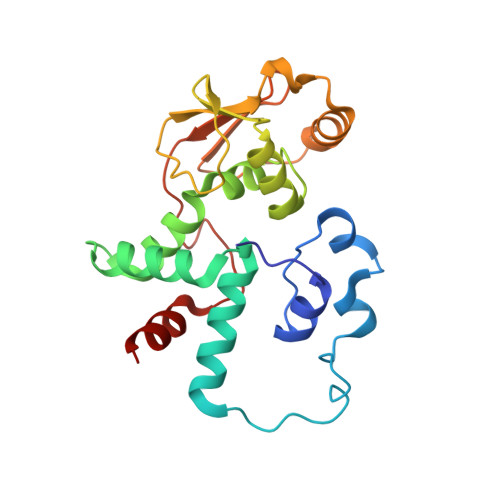
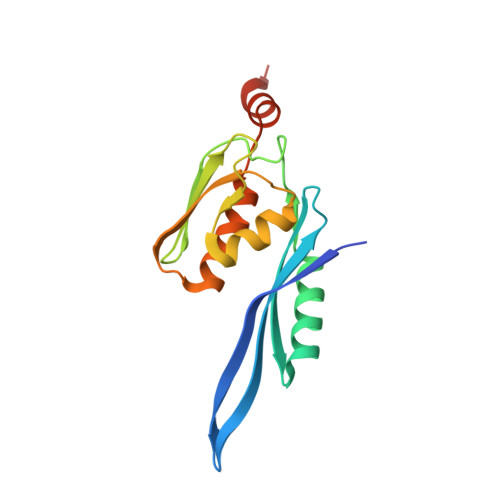
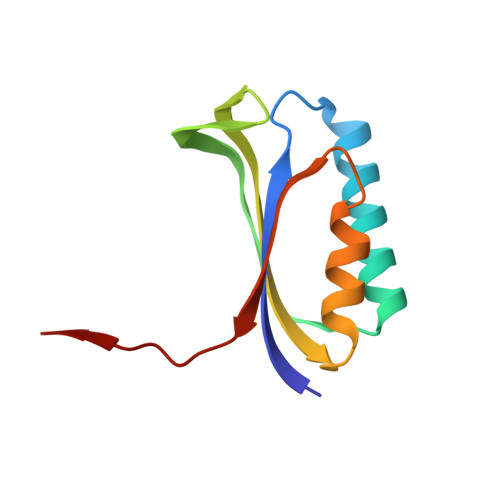
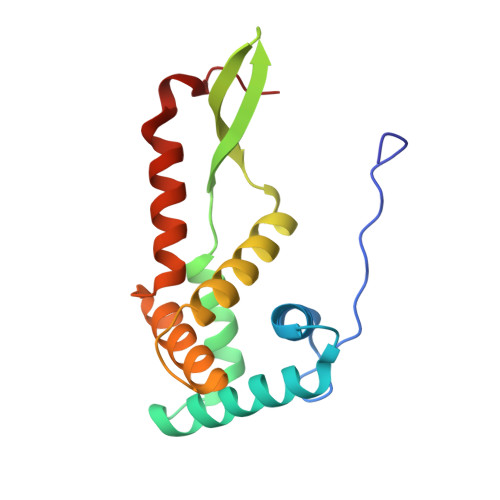
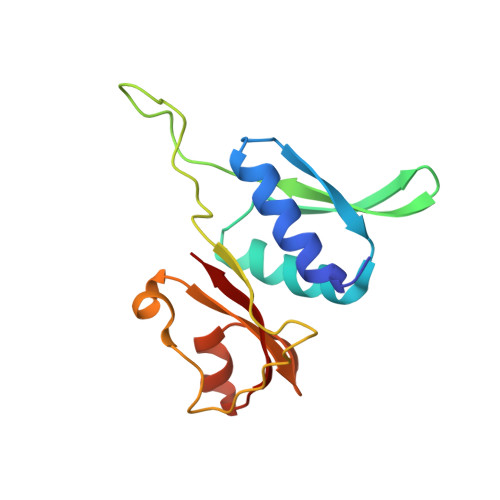
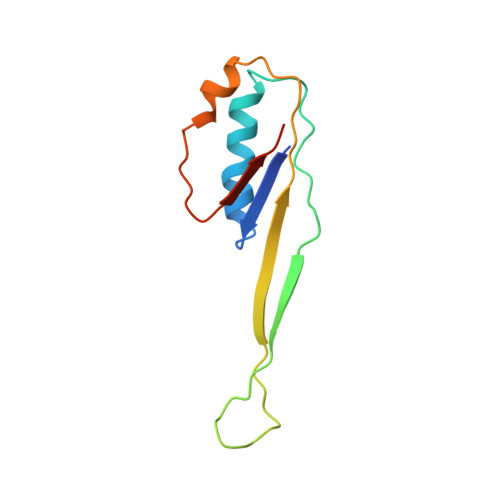
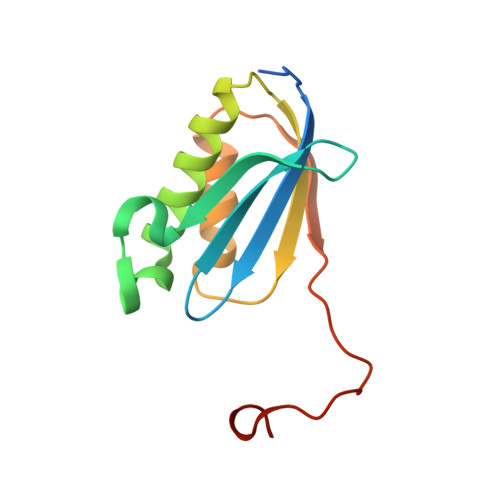
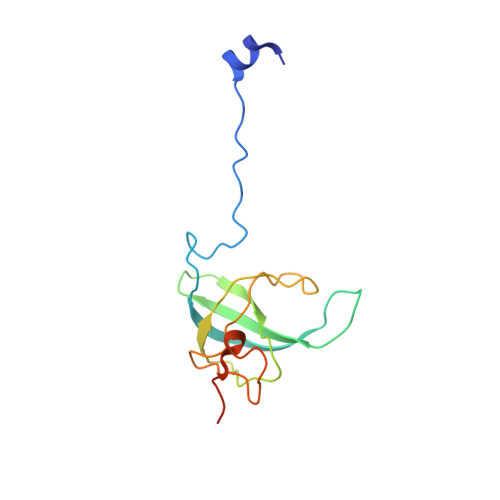
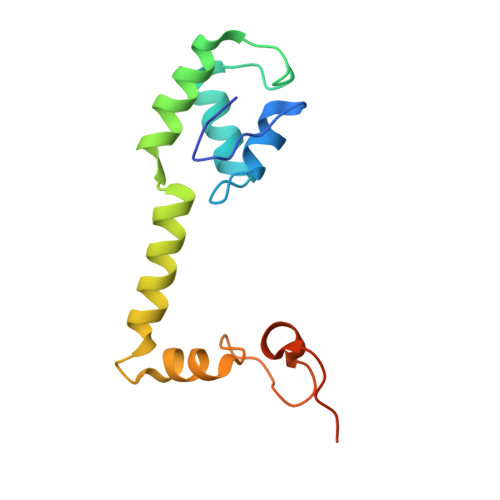
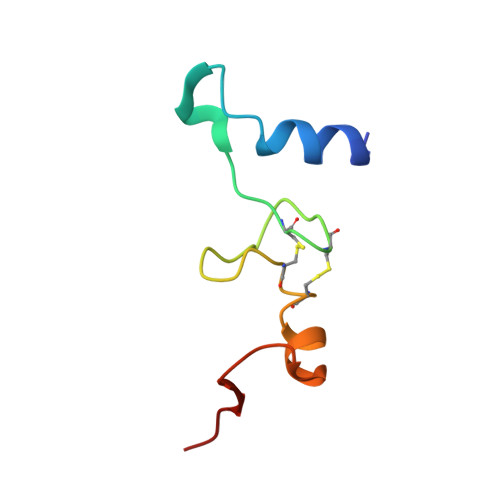
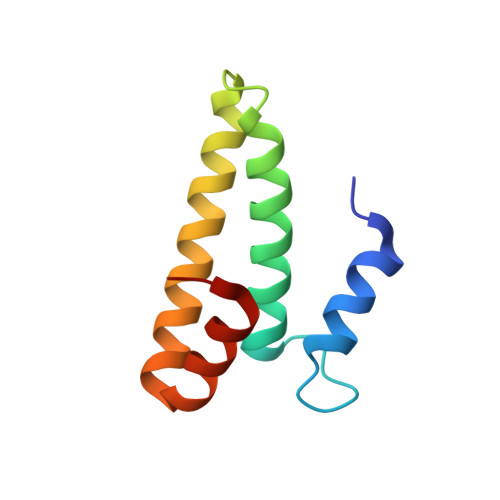
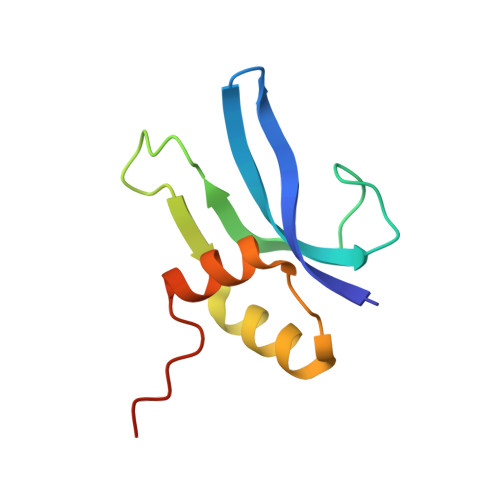
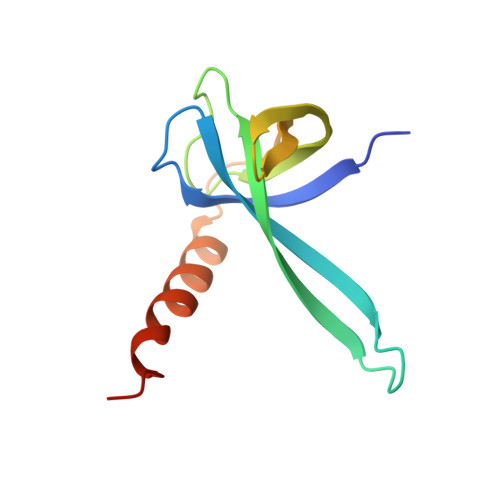
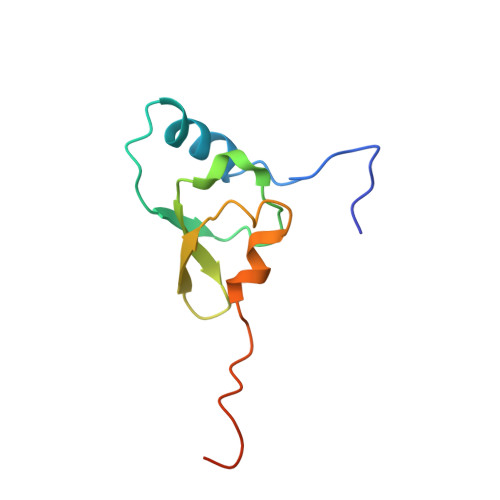
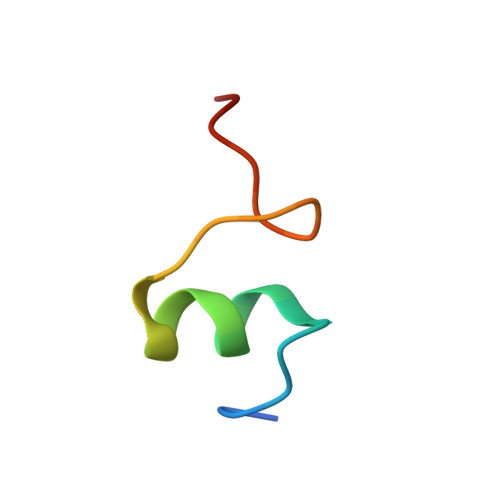
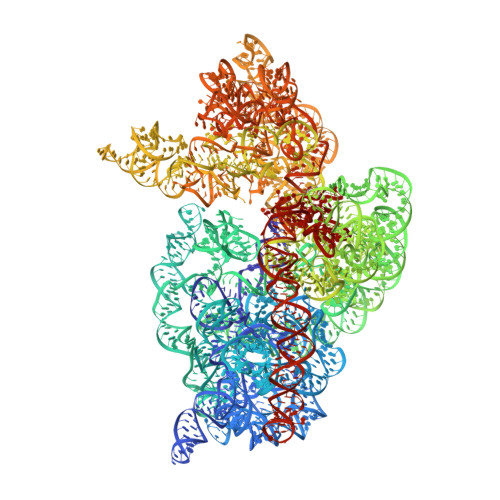
Monomeric YoeB toxin retains RNase activity but adopts an obligate dimeric form for thermal stability.
Pavelich, I.J., Maehigashi, T., Hoffer, E.D., Ruangprasert, A., Miles, S.J., Dunham, C.M.(2019) Nucleic Acids Res 47: 10400-10413
- PubMed: 31501867 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz760
- Primary Citation Related Structures:
6NY6, 6OTR, 6OXA, 6OXI - PubMed Abstract:
Chromosomally-encoded toxin-antitoxin complexes are ubiquitous in bacteria and regulate growth through the release of the toxin component typically in a stress-dependent manner. Type II ribosome-dependent toxins adopt a RelE-family RNase fold and inhibit translation by degrading mRNAs while bound to the ribosome. Here, we present biochemical and structural studies of the Escherichia coli YoeB toxin interacting with both a UAA stop and an AAU sense codon in pre- and post-mRNA cleavage states to provide insights into possible mRNA substrate selection. Both mRNAs undergo minimal changes during the cleavage event in contrast to type II ribosome-dependent RelE toxin. Further, the 16S rRNA decoding site nucleotides that monitor the mRNA in the aminoacyl(A) site adopt different orientations depending upon which toxin is present. Although YoeB is a RelE family member, it is the sole ribosome-dependent toxin that is dimeric. We show that engineered monomeric YoeB is active against mRNAs bound to both the small and large subunit. However, the stability of monomeric YoeB is reduced ∼20°C, consistent with potential YoeB activation during heat shock in E. coli as previously demonstrated. These data provide a molecular basis for the ability of YoeB to function in response to thermal stress.
- Department of Biochemistry, Emory University School of Medicine, Atlanta, GA 30322, USA.
Organizational Affiliation: