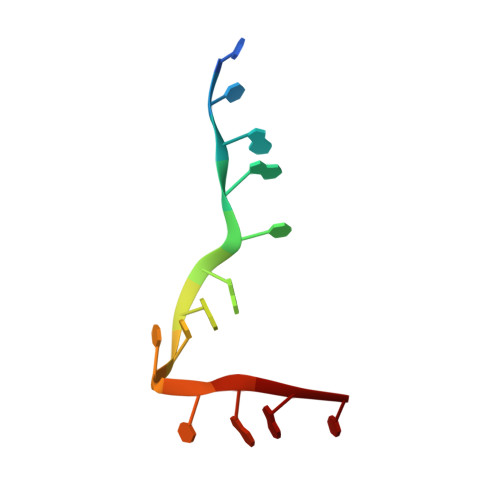
Distinct RNA-binding modules in a single PUF protein cooperate to determine RNA specificity.
Qiu, C., Dutcher, R.C., Porter, D.F., Arava, Y., Wickens, M., Hall, T.M.T.(2019) Nucleic Acids Res 47: 8770-8784
- PubMed: 31294800 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz583
- Primary Citation Related Structures:
6NWW, 6NX5, 6NY5 - PubMed Abstract:
PUF proteins, named for Drosophila Pumilio (PUM) and Caenorhabditis elegans fem-3-binding factor (FBF), recognize specific sequences in the mRNAs they bind and control. RNA binding by classical PUF proteins is mediated by a characteristic PUM homology domain (PUM-HD). The Puf1 and Puf2 proteins possess a distinct architecture and comprise a highly conserved subfamily among fungal species. Puf1/Puf2 proteins contain two types of RNA-binding domain: a divergent PUM-HD and an RNA recognition motif (RRM). They recognize RNAs containing UAAU motifs, often in clusters. Here, we report a crystal structure of the PUM-HD of a fungal Puf1 in complex with a dual UAAU motif RNA. Each of the two UAAU tetranucleotides are bound by a Puf1 PUM-HD forming a 2:1 protein-to-RNA complex. We also determined crystal structures of the Puf1 RRM domain that identified a dimerization interface. The PUM-HD and RRM domains act in concert to determine RNA-binding specificity: the PUM-HD dictates binding to UAAU, and dimerization of the RRM domain favors binding to dual UAAU motifs rather than a single UAAU. Cooperative action of the RRM and PUM-HD identifies a new mechanism by which multiple RNA-binding modules in a single protein collaborate to create a unique RNA-binding specificity.
- Epigenetics and Stem Cell Biology Laboratory, National Institute of Environmental Health Sciences, National Institutes of Health, Research Triangle Park, NC 27709, USA.
Organizational Affiliation: