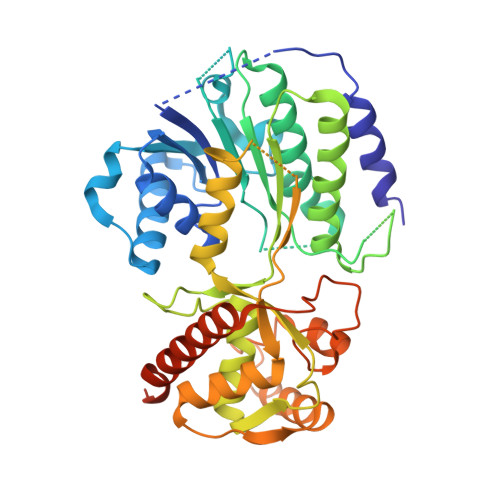
Fungal indole alkaloid biogenesis through evolution of a bifunctional reductase/Diels-Alderase.
Dan, Q., Newmister, S.A., Klas, K.R., Fraley, A.E., McAfoos, T.J., Somoza, A.D., Sunderhaus, J.D., Ye, Y., Shende, V.V., Yu, F., Sanders, J.N., Brown, W.C., Zhao, L., Paton, R.S., Houk, K.N., Smith, J.L., Sherman, D.H., Williams, R.M.(2019) Nat Chem 11: 972-980
- PubMed: 31548667 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41557-019-0326-6
- Primary Citation Related Structures:
6NKH, 6NKI, 6NKK, 6NKM - PubMed Abstract:
Prenylated indole alkaloids such as the calmodulin-inhibitory malbrancheamides and anthelmintic paraherquamides possess great structural diversity and pharmaceutical utility. Here, we report complete elucidation of the malbrancheamide biosynthetic pathway accomplished through complementary approaches. These include a biomimetic total synthesis to access the natural alkaloid and biosynthetic intermediates in racemic form and in vitro enzymatic reconstitution to provide access to the natural antipode (+)-malbrancheamide. Reductive cleavage of an L-Pro-L-Trp dipeptide from the MalG non-ribosomal peptide synthetase (NRPS) followed by reverse prenylation and a cascade of post-NRPS reactions culminates in an intramolecular [4+2] hetero-Diels-Alder (IMDA) cyclization to furnish the bicyclo[2.2.2]diazaoctane scaffold. Enzymatic assembly of optically pure (+)-premalbrancheamide involves an unexpected zwitterionic intermediate where MalC catalyses enantioselective cycloaddition as a bifunctional NADPH-dependent reductase/Diels-Alderase. The crystal structures of substrate and product complexes together with site-directed mutagenesis and molecular dynamics simulations demonstrate how MalC and PhqE (its homologue from the paraherquamide pathway) catalyse diastereo- and enantioselective cyclization in the construction of this important class of secondary metabolites.
- Life Sciences Institute, University of Michigan, Ann Arbor, MI, USA.
Organizational Affiliation: