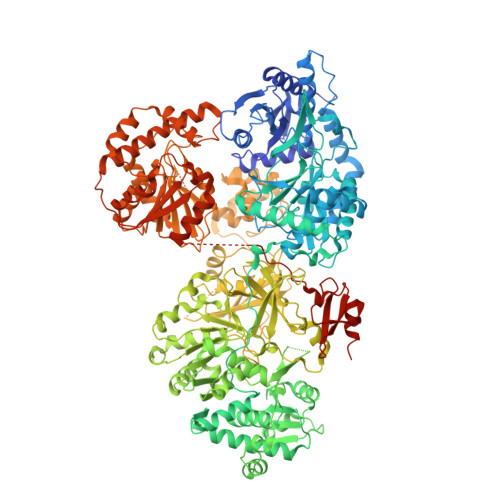
The structural basis of N-acyl-alpha-amino-beta-lactone formation catalyzed by a nonribosomal peptide synthetase.
Kreitler, D.F., Gemmell, E.M., Schaffer, J.E., Wencewicz, T.A., Gulick, A.M.(2019) Nat Commun 10: 3432-3432
- PubMed: 31366889 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-11383-7
- Primary Citation Related Structures:
6N8E - PubMed Abstract:
Nonribosomal peptide synthetases produce diverse natural products using a multidomain architecture where the growing peptide, attached to an integrated carrier domain, is delivered to neighboring catalytic domains for bond formation and modification. Investigation of these systems can lead to the discovery of new structures, unusual biosynthetic transformations, and to the engineering of catalysts for generating new products. The antimicrobial β-lactone obafluorin is produced nonribosomally from dihydroxybenzoic acid and a β-hydroxy amino acid that cyclizes into the β-lactone during product release. Here we report the structure of the nonribosomal peptide synthetase ObiF1, highlighting the structure of the β-lactone-producing thioesterase domain and an interaction between the C-terminal MbtH-like domain with an upstream adenylation domain. Biochemical assays examine catalytic promiscuity, provide mechanistic insight, and demonstrate utility for generating obafluorin analogs. These results advance our understanding of the structural cycle of nonribosomal peptide synthetases and provide insights into the production of β-lactone natural products.
- Department of Structural Biology, Jacobs School of Medicine & Biomedical Sciences at the University at Buffalo, 955 Main Street, Buffalo, NY, 14203, USA.
Organizational Affiliation: