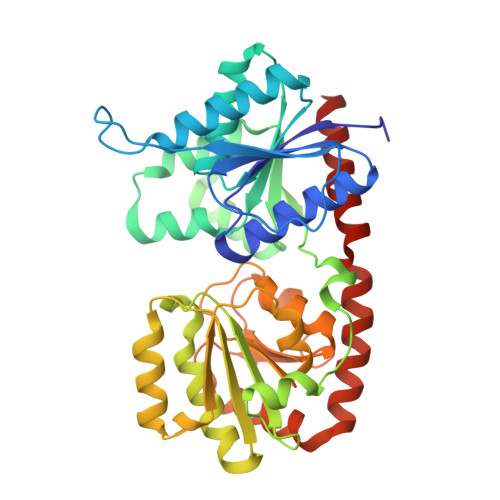
A structural and functional analysis of the glycosyltransferase BshA from Staphylococcus aureus: Insights into the reaction mechanism and regulation of bacillithiol production.
Royer, C.J., Cook, P.D.(2019) Protein Sci 28: 1083-1094
- PubMed: 30968475 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3617
- Primary Citation Related Structures:
6D9T, 6N1X - PubMed Abstract:
Bacillithiol is a glucosamine-derived antioxidant found in several pathogenic Gram-positive bacteria. The compound is involved in maintaining the appropriate redox state within the cell as well as detoxifying foreign agents like the antibiotic fosfomycin. Bacillithiol is produced via the action of three enzymes, including BshA, a retaining GT-B glycosyltransferase that utilizes UDP-N-acetylglucosamine and l-malate to produce N-acetylglucosaminyl-malate. Recent studies suggest that retaining GT-B glycosyltransferases like BshA utilize a substrate-assisted mechanism that goes through an S N i-like transition state. In a previous study, we relied on X-ray crystallography as well as computational simulations to hypothesize the manner in which substrates would bind the enzyme, but several questions about substrate binding and the role of one of the amino acid residues persisted. Another study demonstrated that BshA might be subject to feedback inhibition by bacillithiol, but this phenomenon was not analyzed further to determine the exact mechanism of inhibition. Here we present X-ray crystallographic structures and steady-state kinetics results that help elucidate both of these issues. Our ligand-bound crystal structures demonstrate that the active site provides an appropriate steric and geometric arrangement of ligands to facilitate the substrate-assisted mechanism. Finally, we show that bacillithiol is competitive for UDP-N-acetylglucosamine with a K i value near 120-130 μM and likely binds within the BshA active site, suggesting that bacillithiol modulates BshA activity via feedback inhibition. The work presented here furthers our understanding of bacillithiol metabolism and can aid in the development of inhibitors to counteract resistance to antibiotics such as fosfomycin.
- Department of Chemistry, Grand Valley State University, Allendale, Michigan.
Organizational Affiliation: