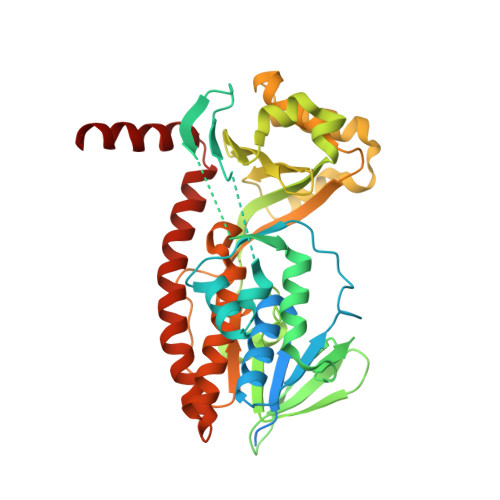
The crystal structure of AbsH3: A putative flavin adenine dinucleotide-dependent reductase in the abyssomicin biosynthesis pathway.
Clinger, J.A., Wang, X., Cai, W., Zhu, Y., Miller, M.D., Zhan, C.G., Van Lanen, S.G., Thorson, J.S., Phillips Jr., G.N.(2020) Proteins
- PubMed: 32852843 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.25994
- Primary Citation Related Structures:
6N04 - PubMed Abstract:
Natural products and natural product-derived compounds have been widely used for pharmaceuticals for many years, and the search for new natural products that may have interesting activity is ongoing. Abyssomicins are natural product molecules that have antibiotic activity via inhibition of the folate synthesis pathway in microbiota. These compounds also appear to undergo a required [4 + 2] cycloaddition in their biosynthetic pathway. Here we report the structure of an flavin adenine dinucleotide-dependent reductase, AbsH3, from the biosynthetic gene cluster of novel abyssomicins found in Streptomyces sp. LC-6-2.
- Department of Biosciences, Rice University, Houston, Texas, USA.
Organizational Affiliation: