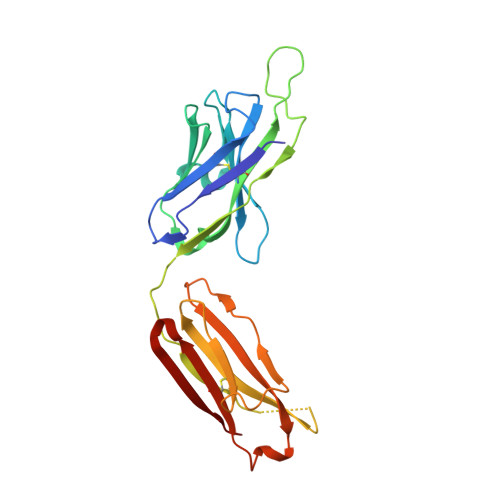
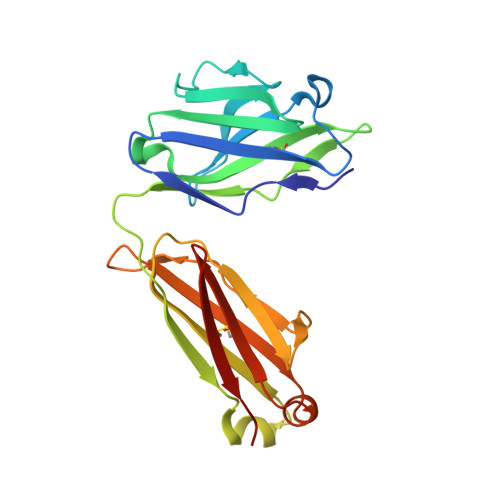
Longitudinal Analysis Reveals Early Development of Three MPER-Directed Neutralizing Antibody Lineages from an HIV-1-Infected Individual.
Krebs, S.J., Kwon, Y.D., Schramm, C.A., Law, W.H., Donofrio, G., Zhou, K.H., Gift, S., Dussupt, V., Georgiev, I.S., Schatzle, S., McDaniel, J.R., Lai, Y.T., Sastry, M., Zhang, B., Jarosinski, M.C., Ransier, A., Chenine, A.L., Asokan, M., Bailer, R.T., Bose, M., Cagigi, A., Cale, E.M., Chuang, G.Y., Darko, S., Driscoll, J.I., Druz, A., Gorman, J., Laboune, F., Louder, M.K., McKee, K., Mendez, L., Moody, M.A., O'Sullivan, A.M., Owen, C., Peng, D., Rawi, R., Sanders-Buell, E., Shen, C.H., Shiakolas, A.R., Stephens, T., Tsybovsky, Y., Tucker, C., Verardi, R., Wang, K., Zhou, J., Zhou, T., Georgiou, G., Alam, S.M., Haynes, B.F., Rolland, M., Matyas, G.R., Polonis, V.R., McDermott, A.B., Douek, D.C., Shapiro, L., Tovanabutra, S., Michael, N.L., Mascola, J.R., Robb, M.L., Kwong, P.D., Doria-Rose, N.A.(2019) Immunity 50: 677-691.e13
- PubMed: 30876875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2019.02.008
- Primary Citation Related Structures:
6MTO, 6MTP, 6MTQ, 6MTR, 6MTS, 6MTT - PubMed Abstract:
Lineage-based vaccine design is an attractive approach for eliciting broadly neutralizing antibodies (bNAbs) against HIV-1. However, most bNAb lineages studied to date have features indicative of unusual recombination and/or development. From an individual in the prospective RV217 cohort, we identified three lineages of bNAbs targeting the membrane-proximal external region (MPER) of the HIV-1 envelope. Antibodies RV217-VRC42.01, -VRC43.01, and -VRC46.01 used distinct modes of recognition and neutralized 96%, 62%, and 30%, respectively, of a 208-strain virus panel. All three lineages had modest levels of somatic hypermutation and normal antibody-loop lengths and were initiated by the founder virus MPER. The broadest lineage, VRC42, was similar to the known bNAb 4E10. A multimeric immunogen based on the founder MPER activated B cells bearing the unmutated common ancestor of VRC42, with modest maturation of early VRC42 intermediates imparting neutralization breadth. These features suggest that VRC42 may be a promising template for lineage-based vaccine design.
- U.S. Military HIV Research Program, WRAIR, Silver Spring, MD, USA; Henry Jackson Foundation, Bethesda, MD, USA.
Organizational Affiliation: