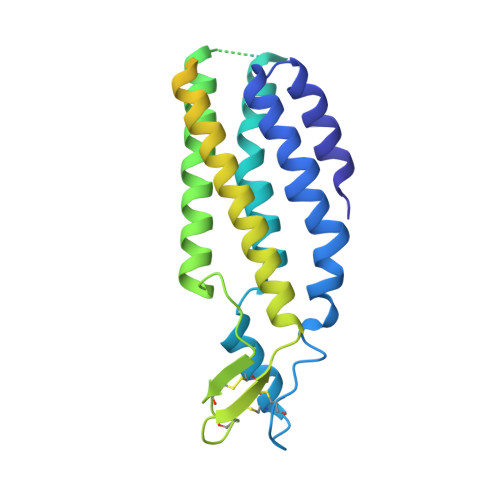
Structure of native lens connexin 46/50 intercellular channels by cryo-EM.
Myers, J.B., Haddad, B.G., O'Neill, S.E., Chorev, D.S., Yoshioka, C.C., Robinson, C.V., Zuckerman, D.M., Reichow, S.L.(2018) Nature 564: 372-377
- PubMed: 30542154 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-018-0786-7
- Primary Citation Related Structures:
6MHQ, 6MHY - PubMed Abstract:
Gap junctions establish direct pathways for cell-to-cell communication through the assembly of twelve connexin subunits that form intercellular channels connecting neighbouring cells. Co-assembly of different connexin isoforms produces channels with unique properties and enables communication across cell types. Here we used single-particle cryo-electron microscopy to investigate the structural basis of connexin co-assembly in native lens gap junction channels composed of connexin 46 and connexin 50 (Cx46/50). We provide the first comparative analysis to connexin 26 (Cx26), which-together with computational studies-elucidates key energetic features governing gap junction permselectivity. Cx46/50 adopts an open-state conformation that is distinct from the Cx26 crystal structure, yet it appears to be stabilized by a conserved set of hydrophobic anchoring residues. 'Hot spots' of genetic mutations linked to hereditary cataract formation map to the core structural-functional elements identified in Cx46/50, suggesting explanations for many of the disease-causing effects.
- Department of Chemistry, Portland State University, Portland, OR, USA.
Organizational Affiliation: