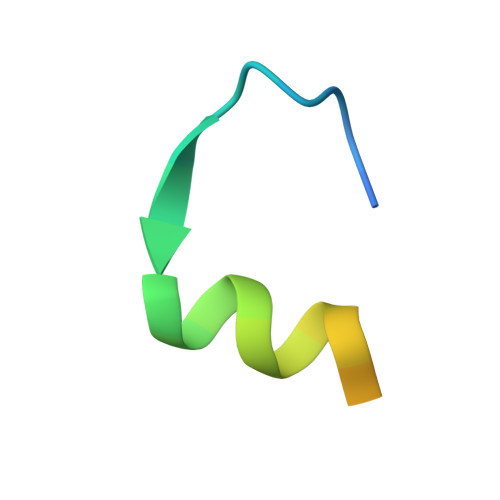
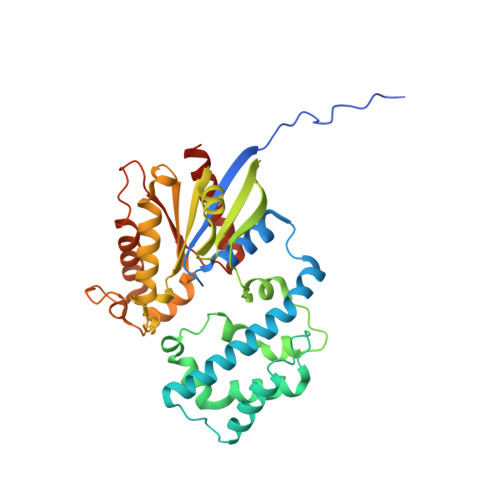
Structural basis for GPCR-independent activation of heterotrimeric Gi proteins.
Kalogriopoulos, N.A., Rees, S.D., Ngo, T., Kopcho, N.J., Ilatovskiy, A.V., Sun, N., Komives, E.A., Chang, G., Ghosh, P., Kufareva, I.(2019) Proc Natl Acad Sci U S A 116: 16394-16403
- PubMed: 31363053 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1906658116
- Primary Citation Related Structures:
6MHE, 6MHF - PubMed Abstract:
Heterotrimeric G proteins are key molecular switches that control cell behavior. The canonical activation of G proteins by agonist-occupied G protein-coupled receptors (GPCRs) has recently been elucidated from the structural perspective. In contrast, the structural basis for GPCR-independent G protein activation by a novel family of guanine-nucleotide exchange modulators (GEMs) remains unknown. Here, we present a 2.0-Å crystal structure of Gαi in complex with the GEM motif of GIV/Girdin. Nucleotide exchange assays, molecular dynamics simulations, and hydrogen-deuterium exchange experiments demonstrate that GEM binding to the conformational switch II causes structural changes that allosterically propagate to the hydrophobic core of the Gαi GTPase domain. Rearrangement of the hydrophobic core appears to be a common mechanism by which GPCRs and GEMs activate G proteins, although with different efficiency. Atomic-level insights presented here will aid structure-based efforts to selectively target the noncanonical G protein activation.
- Department of Medicine, University of California San Diego, La Jolla, CA 92093.
Organizational Affiliation: