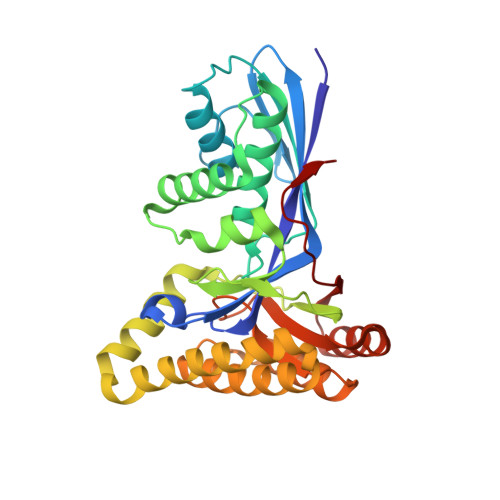
Structural insight into substrate and product binding in an archaeal mevalonate kinase.
Miller, B.R., Kung, Y.(2018) PLoS One 13: e0208419-e0208419
- PubMed: 30521590 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0208419
- Primary Citation Related Structures:
6MDE, 6MDF - PubMed Abstract:
Mevalonate kinase (MK) is a key enzyme of the mevalonate pathway, which produces the biosynthetic precursors for steroids, including cholesterol, and isoprenoids, the largest class of natural products. Currently available crystal structures of MK from different organisms depict the enzyme in its unbound, substrate-bound, and inhibitor-bound forms; however, until now no structure has yet been determined of MK bound to its product, 5-phosphomevalonate. Here, we present crystal structures of mevalonate-bound and 5-phosphomevalonate-bound MK from Methanosarcina mazei (MmMK), a methanogenic archaeon. In contrast to the prior structure of a eukaryotic MK bound with mevalonate, we find a striking lack of direct interactions between this archaeal MK and its substrate. Further, these two MmMK structures join the prior structure of the apoenzyme to complete the first suite of structural snapshots that depict unbound, substrate-bound, and product-bound forms of the same MK. With this collection of structures, we now provide additional insight into the catalytic mechanism of this biologically essential enzyme.
- Department of Chemistry, Bryn Mawr College, Bryn Mawr, PA, United States of America.
Organizational Affiliation: