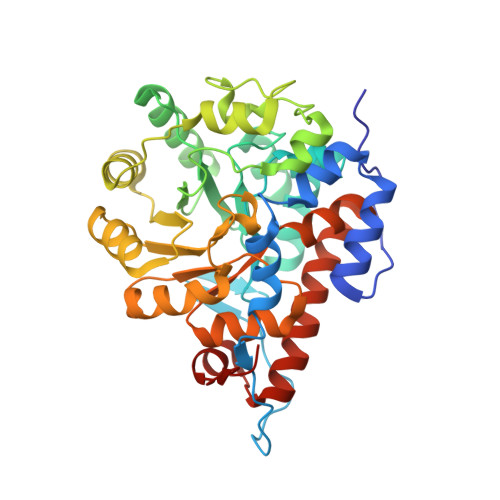
Structure of lactate oxidase from Enterococcus hirae revealed new aspects of active site loop function: Product-inhibition mechanism and oxygen gatekeeper
Hiraka, K., Yoshida, H., Tsugawa, W., Asano, R., La Belle, J.T., Ikebukuro, K., Sode, K.(2022) Protein Sci 31