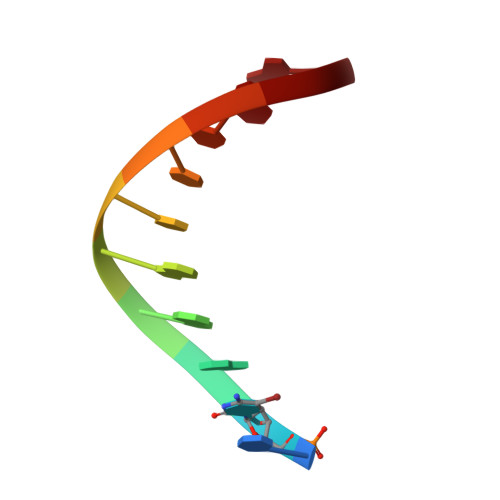
X-ray Crystal Structure of a Cyclic-PIP-DNA Complex in the Reverse-Binding Orientation.
Abe, K., Hirose, Y., Eki, H., Takeda, K., Bando, T., Endo, M., Sugiyama, H.(2020) J Am Chem Soc 142: 10544-10549
- PubMed: 32401492 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.0c03972
- Primary Citation Related Structures:
6M5B - PubMed Abstract:
Elucidation of the details of the associating mode is one of the major concerns for the precise design of DNA-binding molecules that are used for gene regulation. Pyrrole-imidazole polyamide (PIP) is a well-established synthetic DNA-binding molecule that has sequence-specificity for duplex DNA. By the design of the sequence of pyrrole, imidazole, and other synthetic units, PIP is bound to the target DNA sequence selectively. Here, we report the X-ray crystal structure of newly synthesized chiral cyclic PIP (cPIP) complexed with DNA at 1.5 Å resolution and reveal that cPIP binds in the reverse orientation in the DNA minor groove. Analysis of the crystal structure revealed that the positions of the hydrogen bonds between the bases and the pyrrole-imidazole moieties of cPIP were similar for both forward- and reverse-binding orientations and that the distortion of the B-form DNA structure caused by cPIP binding was also similar for both orientations. We further found that new hydrogen bonds formed between the amino groups on the γ-turn units and DNA at both ends of the cPIP molecule. Additionally, by comparing the reverse PIP orientation with the forward orientation, we could clarify that the cause of the preference toward the reverse orientation in the S -form cPIP as used in this study is the overall conformation of the cPIP-DNA complex, particularly the configuration of hydrogen bonds. These results thus provide an explanation for the different stereoselectivity of cPIP binding in the minor groove.
- Department of Chemistry, Graduate School of Science, Kyoto University, Kitashirakawa-oiwakecho, Sakyo-ku, Kyoto 606-8502, Japan.
Organizational Affiliation: