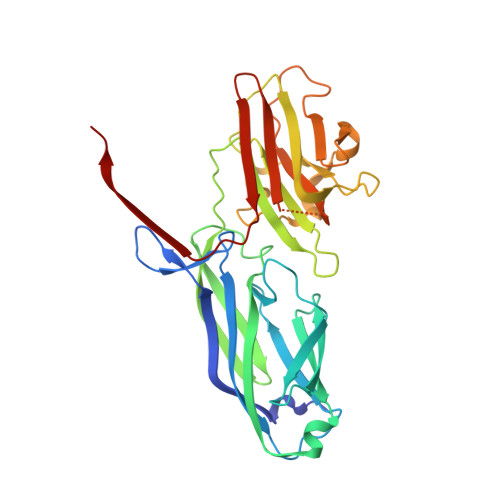
Structural Basis ofStaphylococcus aureusSurface Protein SdrC.
Pi, Y., Chen, W., Ji, Q.(2020) Biochemistry 59: 1465-1469
- PubMed: 32250096 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.0c00124
- Primary Citation Related Structures:
6LXH, 6LXS - PubMed Abstract:
Staphylococcus aureus surface proteins play important roles in host tissue colonization, biofilm formation, and bacterial virulence and are thus essential for successful host infections. The surface protein SdrC from S. aureus induces bacterial biofilm formation via an intermolecular homophilic interaction of its N2 domains. However, the molecular mechanism of how the homophilic interaction is achieved is unknown. Here, we report two crystal structures of SdrC N2N3 domains, revealing two possible homophilic interaction mechanisms: Ca 2+ -mediated intermolecular metal chelation of N2 domains and intermolecular interaction of N2 and N3 domains. Given the unnecessary role of the N3 domain in the induction of biofilm formation, the N2 domain-mediated metal chelation mechanism is likely the mechanism that facilitates SdrC homophilic interaction. Mutation of key Ca 2+ -chelating residues differentially reduced the level of protein dimer formation, further supporting the key role of metal chelation in the N2 domain interaction. Together, these results reveal the possible mechanism of the homophilic interaction of SdrC N2 domains and pave the way for the rational development of new strategies against this mechanism.
- School of Physical Science and Technology, ShanghaiTech University, Shanghai 201210, China.
Organizational Affiliation: