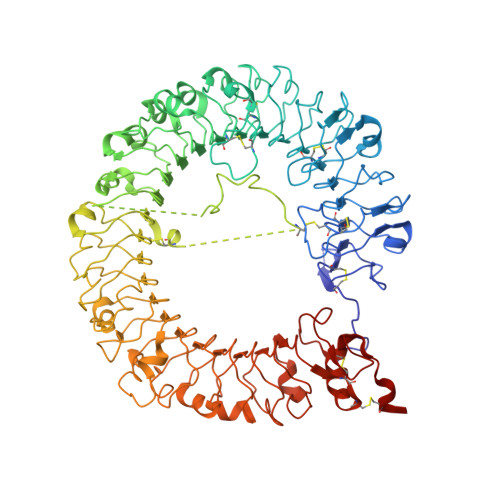
Structural analysis reveals TLR7 dynamics underlying antagonism.
Tojo, S., Zhang, Z., Matsui, H., Tahara, M., Ikeguchi, M., Kochi, M., Kamada, M., Shigematsu, H., Tsutsumi, A., Adachi, N., Shibata, T., Yamamoto, M., Kikkawa, M., Senda, T., Isobe, Y., Ohto, U., Shimizu, T.(2020) Nat Commun 11: 5204-5204
- PubMed: 33060576 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-19025-z
- Primary Citation Related Structures:
6LVX, 6LVY, 6LVZ, 6LW0, 6LW1 - PubMed Abstract:
Toll-like receptor 7 (TLR7) recognizes both microbial and endogenous RNAs and nucleosides. Aberrant activation of TLR7 has been implicated in several autoimmune diseases including systemic lupus erythematosus (SLE). Here, by modifying potent TLR7 agonists, we develop a series of TLR7-specific antagonists as promising therapeutic agents for SLE. These compounds protect mice against lethal autoimmunity. Combining crystallography and cryo-electron microscopy, we identify the open conformation of the receptor and reveal the structural equilibrium between open and closed conformations that underlies TLR7 antagonism, as well as the detailed mechanism by which TLR7-specific antagonists bind to their binding pocket in TLR7. Our work provides small-molecule TLR7-specific antagonists and suggests the TLR7-targeting strategy for treating autoimmune diseases.
- Sumitomo Dainippon Pharma Co., Ltd., 3-1-98 Kasugade-naka, Konohana-ku, Osaka, 554-0022, Japan. shingo-tojo@ds-pharma.co.jp.
Organizational Affiliation: