The Combination of CD8 alpha alpha and Peptide-MHC-I in a Face-to-Face Mode Promotes Chicken gamma delta T Cells Response.
Liu, Y., Chen, R., Liang, R., Sun, B., Wu, Y., Zhang, L., Kaufman, J., Xia, C.(2020) Front Immunol 11: 605085-605085
- PubMed: 33329601 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fimmu.2020.605085
- Primary Citation Related Structures:
6LHF, 6LHG, 6LHH - PubMed Abstract:
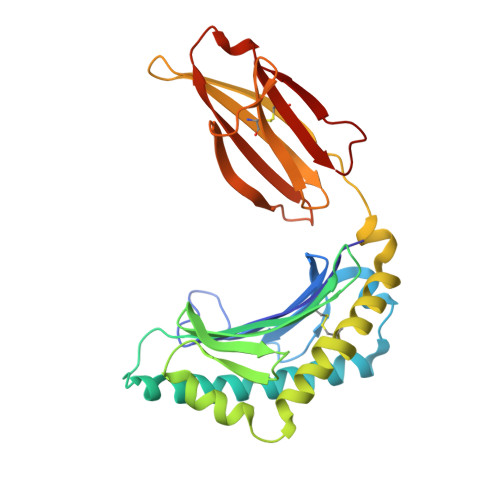
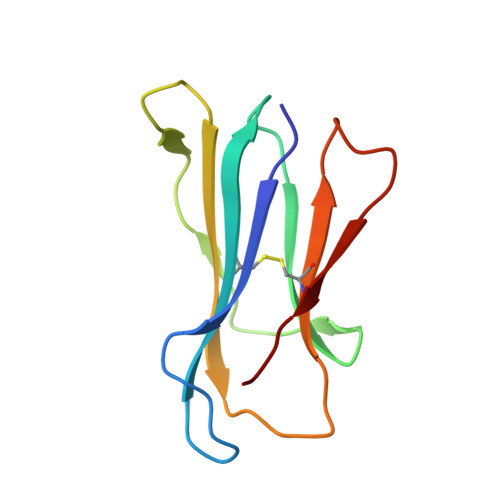
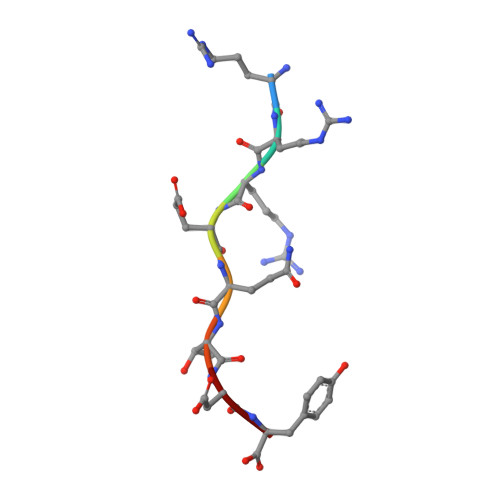
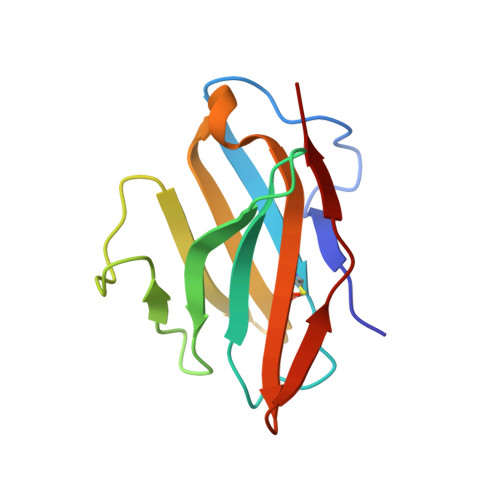
The CD8αα homodimer is crucial to both thymic T cell selection and the antigen recognition of cytotoxic T cells. The CD8-pMHC-I interaction can enhance CTL immunity via stabilizing the TCR-pMHC-I interaction and optimizing the cross-reactivity and Ag sensitivity of CD8 + T cells at various stages of development. To date, only human and mouse CD8-pMHC-I complexes have been determined. Here, we resolved the pBF2*1501 complex and the cCD8αα/pBF2*1501 and cCD8αα/pBF2*0401 complexes in nonmammals for the first time. Remarkably, cCD8αα/pBF2*1501 and the cCD8αα/pBF2*0401 complex both exhibited two binding modes, including an "antibody-like" mode similar to that of the known mammal CD8/pMHC-I complexes and a "face-to-face" mode that has been observed only in chickens to date. Compared to the "antibody-like" mode, the "face-to-face" binding mode changes the binding orientation of the cCD8αα homodimer to pMHC-I, which might facilitate abundant γδT cells to bind diverse peptides presented by limited BF2 alleles in chicken. Moreover, the forces involving in the interaction of cCD8αα/pBF2*1501 and the cCD8αα/pBF2*0401 are different in this two binding model, which might change the strength of the CD8-pMHC-I interaction, amplifying T cell cross-reactivity in chickens. The coreceptor CD8αα of TCR has evolved two peptide-MHC-I binding patterns in chickens, which might enhance the T cell response to major or emerging pathogens, including chicken-derived pathogens that are relevant to human health, such as high-pathogenicity influenza viruses.
- Department of Microbiology and Immunology, College of Veterinary Medicine, China Agricultural University, Beijing, China.
Organizational Affiliation: