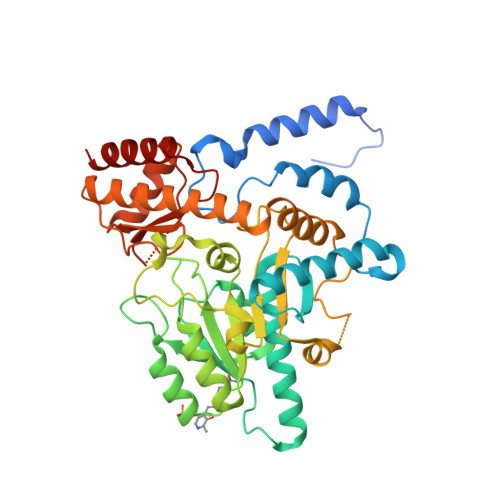
Structural insights into the mechanism of internal aldimine formation and catalytic loop dynamics in an archaeal Group II decarboxylase.
Chellam Gayathri, S., Manoj, N.(2019) J Struct Biol 208: 137-151
- PubMed: 31445086 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2019.08.009
- Primary Citation Related Structures:
6JY1, 6LDR, 6LDS, 6LDT - PubMed Abstract:
Formation of the internal aldimine (LLP) is the first regulatory step that activates pyridoxal 5'-phosphate (PLP) dependent enzymes. The process involves a nucleophilic attack on PLP by an active site Lys residue, followed by proton transfers resulting in a carbinolamine (CBA) intermediate that undergoes dehydration to form the aldimine. Despite a general understanding of the pathway, the structural basis of the mechanistic roles of specific residues in each of these steps is unclear. Here we determined the crystal structure of the LLP form (holo-form) of a Group II PLP-dependent decarboxylase from Methanocaldococcus jannaschii (MjDC) at 1.7 Å resolution. By comparing the crystal structure of MjDC in the LLP form with that of the pyridoxal-P (non-covalently bound aldehyde) form, we demonstrate structural evidence for a water-mediated mechanism of LLP formation. A conserved extended hydrogen-bonding network around PLP coupled to the pyridinyl nitrogen influences activation and catalysis by affecting the electronic configuration of PLP. Furthermore, the two cofactor bound forms revealed open and closed conformations of the catalytic loop (CL) in the absence of a ligand, supporting a hypothesis for a regulatory link between LLP formation and CL dynamics. The evidence suggests that activation of Group II decarboxylases involves a complex interplay of interactions between the electronic states of PLP, the active site micro-environment and CL dynamics.
- Department of Biotechnology, Bhupat and Jyoti Mehta School of Biosciences, Indian Institute of Technology Madras, Chennai 600036, India.
Organizational Affiliation: