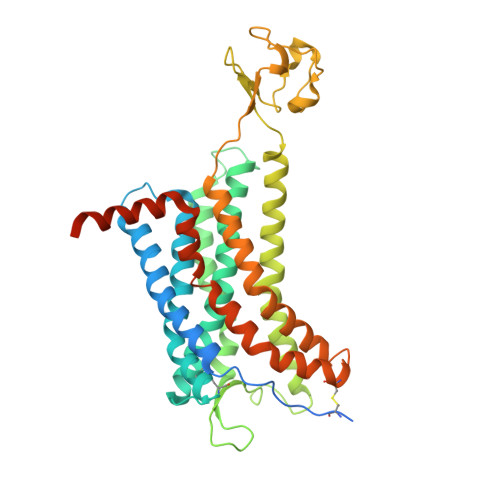
Structure-guided discovery of a single-domain antibody agonist against human apelin receptor.
Ma, Y., Ding, Y., Song, X., Ma, X., Li, X., Zhang, N., Song, Y., Sun, Y., Shen, Y., Zhong, W., Hu, L.A., Ma, Y., Zhang, M.Y.(2020) Sci Adv 6: eaax7379-eaax7379
- PubMed: 31998837 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aax7379
- Primary Citation Related Structures:
6KNM - PubMed Abstract:
Developing antibody agonists targeting the human apelin receptor (APJ) is a promising therapeutic approach for the treatment of chronic heart failure. Here, we report the structure-guided discovery of a single-domain antibody (sdAb) agonist JN241-9, based on the cocrystal structure of APJ with an sdAb antagonist JN241, the first cocrystal structure of a class A G protein-coupled receptor (GPCR) with a functional antibody. As revealed by the structure, JN241 binds to the extracellular side of APJ, makes critical contacts with the second extracellular loop, and inserts the CDR3 into the ligand-binding pocket. We converted JN241 into a full agonist JN241-9 by inserting a tyrosine into the CDR3. Modeling and molecular dynamics simulation shed light on JN241-9-stimulated receptor activation, providing structural insights for finding agonistic antibodies against class A GPCRs.
- Amgen Discovery Research, Amgen Asia R&D Center, Amgen Biopharmaceutical R&D (Shanghai) Co. Ltd., 13th Floor, Building No. 2, 4560 Jinke Road, Zhangjiang, Shanghai 201210, China.
Organizational Affiliation: