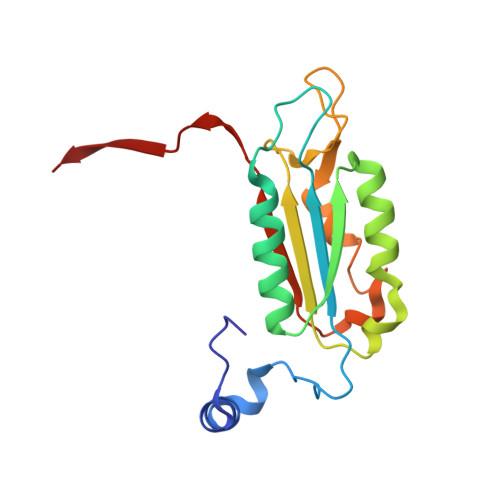
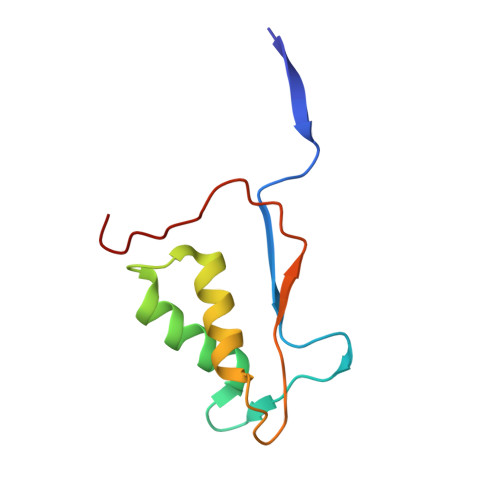
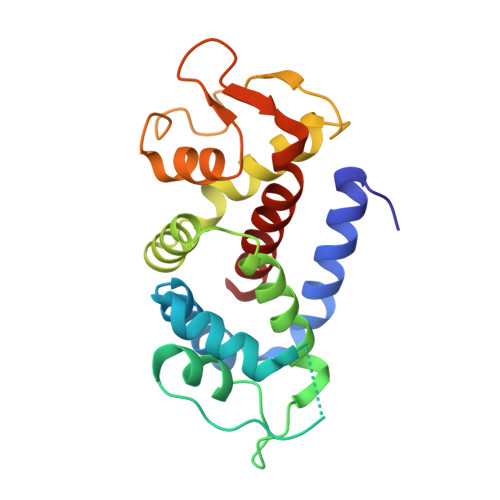
Structural Mechanism for GSDMD Targeting by Autoprocessed Caspases in Pyroptosis.
Wang, K., Sun, Q., Zhong, X., Zeng, M., Zeng, H., Shi, X., Li, Z., Wang, Y., Zhao, Q., Shao, F., Ding, J.(2020) Cell 180: 941
- PubMed: 32109412 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2020.02.002
- Primary Citation Related Structures:
6KMT, 6KMU, 6KMV, 6KMZ, 6KN0, 6KN1 - PubMed Abstract:
The pyroptosis execution protein GSDMD is cleaved by inflammasome-activated caspase-1 and LPS-activated caspase-11/4/5. The cleavage unmasks the pore-forming domain from GSDMD-C-terminal domain. How the caspases recognize GSDMD and its connection with caspase activation are unknown. Here, we show site-specific caspase-4/11 autoprocessing, generating a p10 product, is required and sufficient for cleaving GSDMD and inducing pyroptosis. The p10-form autoprocessed caspase-4/11 binds the GSDMD-C domain with a high affinity. Structural comparison of autoprocessed and unprocessed capase-11 identifies a β sheet induced by the autoprocessing. In caspase-4/11-GSDMD-C complex crystal structures, the β sheet organizes a hydrophobic GSDMD-binding interface that is only possible for p10-form caspase-4/11. The binding promotes dimerization-mediated caspase activation, rendering a cleavage independently of the cleavage-site tetrapeptide sequence. Crystal structure of caspase-1-GSDMD-C complex shows a similar GSDMD-recognition mode. Our study reveals an unprecedented substrate-targeting mechanism for caspases. The hydrophobic interface suggests an additional space for developing inhibitors specific for pyroptotic caspases.
- Peking University-Tsinghua University-National Institute of Biological Sciences Joint Graduate Program, National Institute of Biological Sciences, 102206 Beijing, China; National Institute of Biological Sciences, Beijing, 102206 Beijing, China.
Organizational Affiliation: