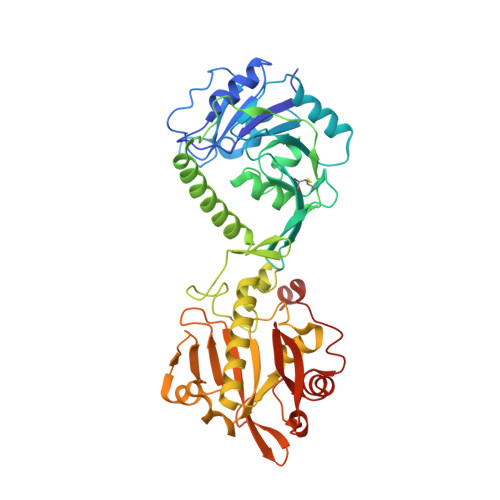
Crystal structures of fukutin-related protein (FKRP), a ribitol-phosphate transferase related to muscular dystrophy.
Kuwabara, N., Imae, R., Manya, H., Tanaka, T., Mizuno, M., Tsumoto, H., Kanagawa, M., Kobayashi, K., Toda, T., Senda, T., Endo, T., Kato, R.(2020) Nat Commun 11: 303-303
- PubMed: 31949166 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-14220-z
- Primary Citation Related Structures:
6KAJ, 6KAK, 6KAL, 6KAM, 6KAN, 6L7S, 6L7T, 6L7U - PubMed Abstract:
α-Dystroglycan (α-DG) is a highly-glycosylated surface membrane protein. Defects in the O-mannosyl glycan of α-DG cause dystroglycanopathy, a group of congenital muscular dystrophies. The core M3 O-mannosyl glycan contains tandem ribitol-phosphate (RboP), a characteristic feature first found in mammals. Fukutin and fukutin-related protein (FKRP), whose mutated genes underlie dystroglycanopathy, sequentially transfer RboP from cytidine diphosphate-ribitol (CDP-Rbo) to form a tandem RboP unit in the core M3 glycan. Here, we report a series of crystal structures of FKRP with and without donor (CDP-Rbo) and/or acceptor [RboP-(phospho-)core M3 peptide] substrates. FKRP has N-terminal stem and C-terminal catalytic domains, and forms a tetramer both in crystal and in solution. In the acceptor complex, the phosphate group of RboP is recognized by the catalytic domain of one subunit, and a phosphate group on O-mannose is recognized by the stem domain of another subunit. Structure-based functional studies confirmed that the dimeric structure is essential for FKRP enzymatic activity.
- Structural Biology Research Center, Institute of Materials Structure Science, High Energy Accelerator Research Organization, Tsukuba, Ibaraki, 305-0801, Japan.
Organizational Affiliation: