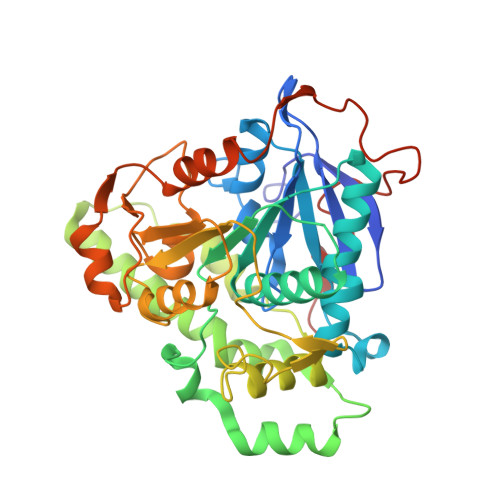
Asymmetric Open-Closed Dimer Mechanism of Polyhydroxyalkanoate Synthase PhaC.
Chek, M.F., Kim, S.Y., Mori, T., Tan, H.T., Sudesh, K., Hakoshima, T.(2020) iScience 23: 101084-101084
- PubMed: 32388399 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2020.101084
- Primary Citation Related Structures:
6K3C - PubMed Abstract:
Biodegradable polyester polyhydroxyalkanoate (PHA) is a promising bioplastic material for industrial use as a replacement for petroleum-based plastics. PHA synthase PhaC forms an active dimer to polymerize acyl moieties from the substrate acyl-coenzyme A (CoA) into PHA polymers. Here we present the crystal structure of the catalytic domain of PhaC from Chromobacterium sp. USM2, bound to CoA. The structure reveals an asymmetric dimer, in which one protomer adopts an open conformation bound to CoA, whereas the other adopts a closed conformation in a CoA-free form. The open conformation is stabilized by the asymmetric dimerization and enables PhaC to accommodate CoA and also to create the product egress path. The bound CoA molecule has its β-mercaptoethanolamine moiety extended into the active site with the terminal SH group close to active center Cys291, enabling formation of the reaction intermediate by acylation of Cys291.
- Structural Biology Laboratory, Nara Institute of Science and Technology, 8916-5 Takayama, Ikoma, Nara 630-0192, Japan.
Organizational Affiliation: