Binding pose 2 of 2-CF3 bound AsqJ complex
Liao, H.J., Chan, N.L.(2020) J Am Chem Soc
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Iron/alpha-ketoglutarate-dependent dioxygenase asqJ | A [auth B] | 327 | Aspergillus nidulans FGSC A4 | Mutation(s): 0 Gene Names: asqJ, AN9227 EC: 1.14 (PDB Primary Data), 1.14.11.81 (UniProt) | 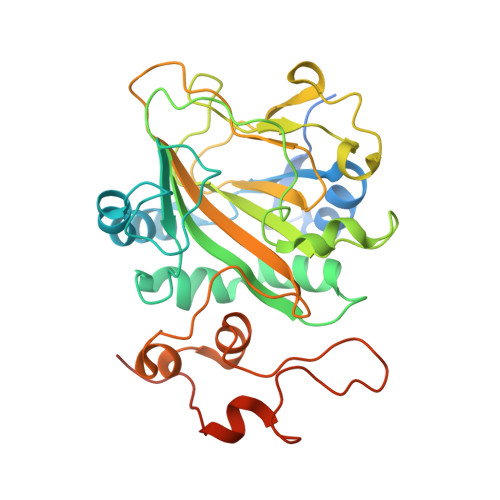 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q5AR53 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CV3 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth B] | (3~{Z})-4-methyl-3-[[4-(trifluoromethyl)phenyl]methylidene]-1~{H}-1,4-benzodiazepine-2,5-dione C18 H13 F3 N2 O2 NVEJUUHKRHQWHW-GDNBJRDFSA-N |  | ||
| SIN Download:Ideal Coordinates CCD File | D [auth B] | SUCCINIC ACID C4 H6 O4 KDYFGRWQOYBRFD-UHFFFAOYSA-N |  | ||
| FE Download:Ideal Coordinates CCD File | B | FE (III) ION Fe VTLYFUHAOXGGBS-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 73.224 | α = 90 |
| b = 118.434 | β = 90 |
| c = 67.336 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data scaling |