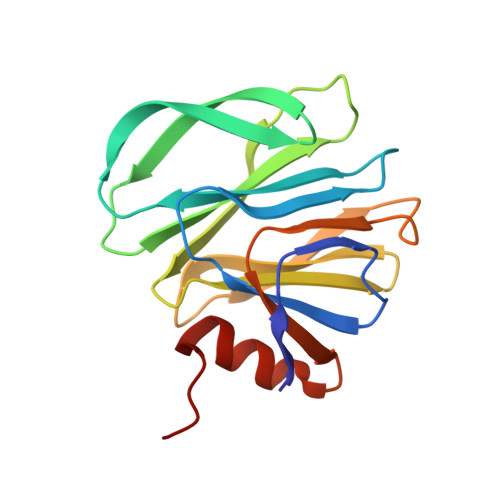
Structural Basis of Glycan Recognition in Globally Predominant Human P[8] Rotavirus.
Sun, X., Dang, L., Li, D., Qi, J., Wang, M., Chai, W., Zhang, Q., Wang, H., Bai, R., Tan, M., Duan, Z.(2020) Virol Sin 35: 156-170
- PubMed: 31620994 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s12250-019-00164-7
- Primary Citation Related Structures:
6K2N, 6K2O - PubMed Abstract:
Rotavirus (RV) causes acute gastroenteritis in infants and children worldwide. Recent studies showed that glycans such as histo-blood group antigens (HBGAs) function as cell attachment factors affecting RV host susceptibility and prevalence. P[8] is the predominant RV genotype in humans, but the structural basis of how P[8] RVs interact with glycan ligands remains elusive. In this study, we characterized the interactions between P[8] VP8*s and glycans which showed that VP8*, the RV glycan binding domain, recognized both mucin core 2 and H type 1 antigens according to the ELISA-based oligosaccharide binding assays. Importantly, we determined the structural basis of P[8] RV-glycans interaction from the crystal structures of a Rotateq P[8] VP8* in complex with core 2 and H type 1 glycans at 1.8 Å and 2.3 Å, respectively, revealing a common binding pocket and similar binding mode. Structural and sequence analysis demonstrated that the glycan binding site is conserved among RVs in the P[II] genogroup, while genotype-specific amino acid variations determined different glycan binding preference. Our data elucidated the detailed structural basis of the interactions between human P[8] RVs and different host glycan factors, shedding light on RV infection, epidemiology, and development of anti-viral agents.
- National Health Commission Key Laboratory for Medical Virology and Viral Diseases, Beijing, 102206, China.
Organizational Affiliation: