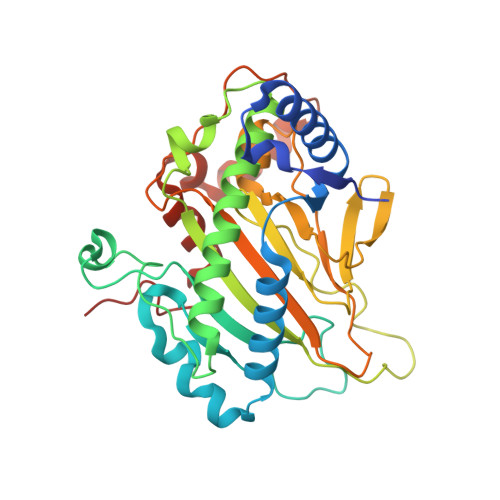
Structural characterization of an isopenicillin N synthase family oxygenase from Pseudomonas aeruginosa PAO1.
Zhang, H., Che, S., Wang, R., Liu, R., Zhang, Q., Bartlam, M.(2019) Biochem Biophys Res Commun 514: 1031-1036
- PubMed: 31097228 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.05.062
- Primary Citation Related Structures:
6JYV - PubMed Abstract:
Isopenicillin N synthase (IPNS) is a nonheme-Fe 2+ -dependent enzyme that mediates a key step in penicillin biosynthesis. It catalyses the conversion of the tripeptide δ-(l-α-aminoadipoyl)-l-cysteine-d-valine (ACV) to isopenicillin N, which is a key precursor to β-lactam antibiotics. The pa4191 gene in Pseudomonas aeruginosa PAO1 has provisionally been annotated as a member of the IPNS family. In this work, we report the crystal structure of PA4191 from P. aeruginosa (PaIPNS hereafter). The 1.65 Å resolution PaIPNS structure forms a jelly roll fold and is confirmed to be a member of the IPNS family based on structural homology. A metal centre within the jelly roll consists of the strictly conserved His201, Asp203 and His257 residues. MicroScale Thermophoresis binding analysis confirms that PaIPNS is a metal-binding protein with a strong preference for iron, but that it does not bind the tripeptide ACV. Structural comparison of PaIPNS with a previously reported IPNS-ACV complex structure reveals a restricted binding pocket that is unable to accommodate ACV.
- College of Life Sciences, Nankai University, Tianjin, 300071, China.
Organizational Affiliation: