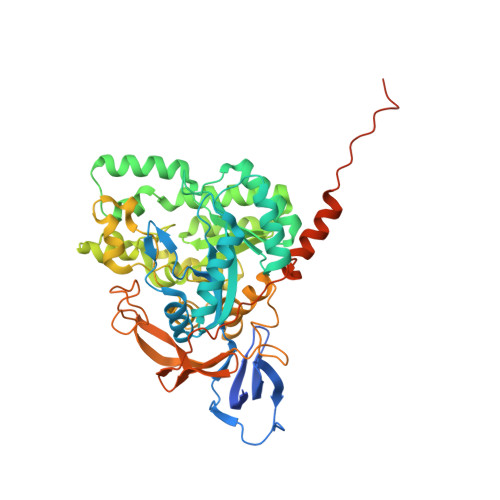
Enhanced carbonyl stress induces irreversible multimerization of CRMP2 in schizophrenia pathogenesis.
Toyoshima, M., Jiang, X., Ogawa, T., Ohnishi, T., Yoshihara, S., Balan, S., Yoshikawa, T., Hirokawa, N.(2019) Life Sci Alliance 2
- PubMed: 31591136 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.201900478
- Primary Citation Related Structures:
6JV9, 6JVB - PubMed Abstract:
Enhanced carbonyl stress underlies a subset of schizophrenia, but its causal effects remain elusive. Here, we elucidated the molecular mechanism underlying the effects of carbonyl stress in iPS cells in which the gene encoding zinc metalloenzyme glyoxalase I ( GLO1 ), a crucial enzyme for the clearance of carbonyl stress, was disrupted. The iPS cells exhibited significant cellular and developmental deficits, and hyper-carbonylation of collapsing response mediator protein 2 (CRMP2). Structural and biochemical analyses revealed an array of multiple carbonylation sites in the functional motifs of CRMP2, particularly D-hook (for dimerization) and T-site (for tetramerization), which are critical for the activity of the CRMP2 tetramer. Interestingly, carbonylated CRMP2 was stacked in the multimer conformation by irreversible cross-linking, resulting in loss of its unique function to bundle microtubules. Thus, the present study revealed that the enhanced carbonyl stress stemmed from the genetic aberrations results in neurodevelopmental deficits through the formation of irreversible dysfunctional multimer of carbonylated CRMP2.
- Laboratory for Molecular Psychiatry, RIKEN Center for Brain Science, Wako, Japan.
Organizational Affiliation: