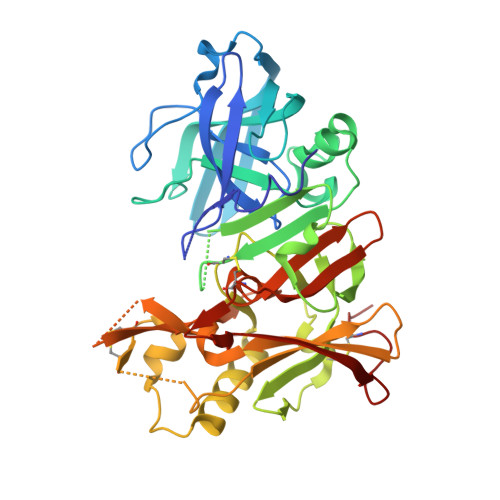
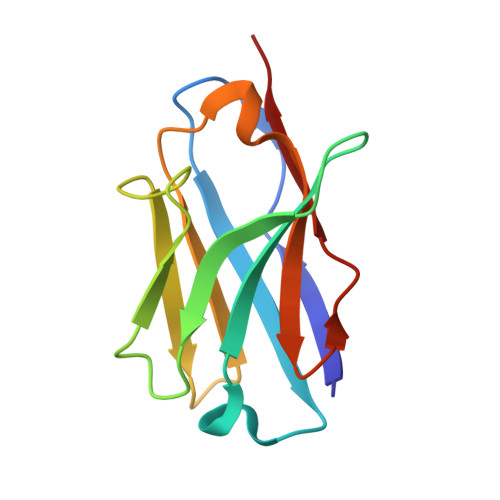
Structure-Based Design of Selective beta-Site Amyloid Precursor Protein Cleaving Enzyme 1 (BACE1) Inhibitors: Targeting the Flap to Gain Selectivity over BACE2.
Fujimoto, K., Matsuoka, E., Asada, N., Tadano, G., Yamamoto, T., Nakahara, K., Fuchino, K., Ito, H., Kanegawa, N., Moechars, D., Gijsen, H.J.M., Kusakabe, K.I.(2019) J Med Chem 62: 5080-5095
- PubMed: 31021626 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b00309
- Primary Citation Related Structures:
6JSE, 6JSF, 6JSG, 6JSN, 6JSZ - PubMed Abstract:
BACE1 inhibitors hold potential as agents in disease-modifying treatment for Alzheimer's disease. BACE2 cleaves the melanocyte protein PMEL in pigment cells of the skin and eye, generating melanin pigments. This role of BACE2 implies that nonselective and chronic inhibition of BACE1 may cause side effects derived from BACE2. Herein, we describe the discovery of potent and selective BACE1 inhibitors using structure-based drug design. We targeted the flap region, where the shape and flexibility differ between these enzymes. Analysis of the cocrystal structures of an initial lead 8 prompted us to incorporate spirocycles followed by its fine-tuning, culminating in highly selective compounds 21 and 22. The structures of 22 bound to BACE1 and BACE2 revealed that a relatively high energetic penalty in the flap of the 22-bound BACE2 structure may cause a loss in BACE2 potency, thereby leading to its high selectivity. These findings and insights should contribute to responding to the challenges in exploring selective BACE1 inhibitors.