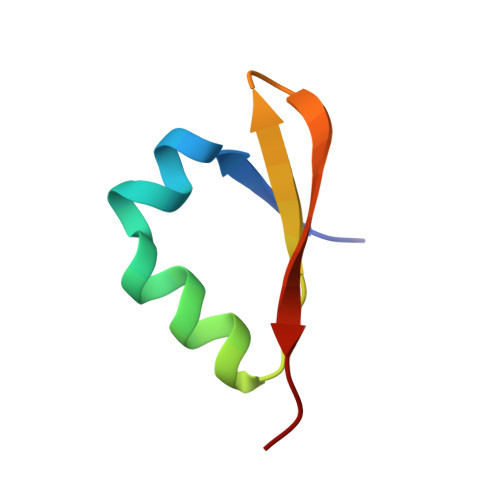
Crystal structure of N-terminal VapB46 antitoxin and interaction analysis of its cognate partners from molecular dynamics simulation.
Roy, M., Saha, R., Singh, B., Yadav, V.K., Bhattacharyya, S., De, S., Das, A.K.(2025) Biochem Biophys Res Commun 783: 152592-152592
- PubMed: 40934556 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2025.152592
- Primary Citation Related Structures:
6JQY - PubMed Abstract:
Toxin-antitoxin (TA) systems are important for persister cell formation in Mycobacterium tuberculosis (Mtb), which aids their survival during stress condition. Antitoxins regulate the TA systems by binding to the promoter-operator region. Among fifty VapBC TA systems in Mtb, few have been characterized, while remaining needs characterisation to understand the VapBC TA mechanism. Here, we report the crystal structure of the N-terminal domain of VapB46 antitoxin at a resolution of 1.64 Å. The protein has a DNA-binding domain that resembles the N-terminal domain of Phd antitoxin with a βααββ fold. Simulation study revealed that VapB46 binds as a dimer at two promoter/operator DNA sites, and tetrameric VapB46 binds to VapC46 toxin via its C-terminal domain. The analysis of the structure and simulation study provides detailed insights into the VapBC46 TA system, paving the way for future drug development targeting this system.
- Department of Bioscience and Biotechnology, Indian Institute of Technology Kharagpur, Kharagpur, 721302, India.
Organizational Affiliation: