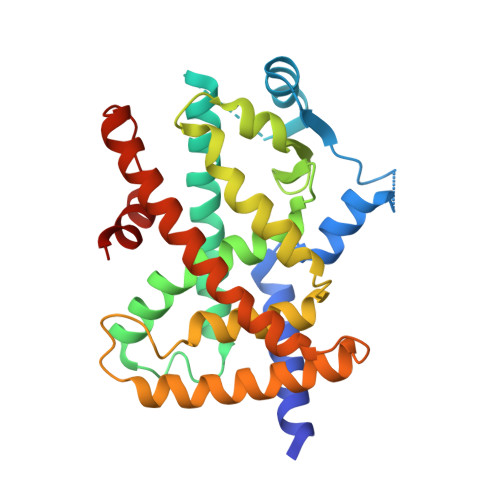
Cyclization Reaction-Based Turn-on Probe for Covalent Labeling of Target Proteins.
Kojima, H., Fujita, Y., Takeuchi, R., Ikebe, Y., Ohashi, N., Yamamoto, K., Itoh, T.(2020) Cell Chem Biol 27: 334
- PubMed: 31991094 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2020.01.006
- Primary Citation Related Structures:
6JEY, 6JEZ, 6JF0 - PubMed Abstract:
Fluorescent molecules have contributed to basic biological research but there are currently only a limited number of probes available for the detection of non-enzymatic proteins. Here, we report turn-on fluorescent probes mediated by conjugate addition and cyclization (TCC probes). These probes react with multiple amino acids and exhibit a 36-fold greater emission intensity after reaction. We analyzed the reactions between TCC probes and nuclear receptors by electrospray ionization mass spectrometry, X-ray crystallography, spectrofluorometry, and fluorescence microscopy. In vitro analysis showed that probes consisting of a protein ligand and TCC could label vitamin D receptor and peroxisome proliferator-activated receptor γ. Moreover, we demonstrated that not only a ligand unit but also a peptide unit can label the target protein in a complex mixture.
- Showa Pharmaceutical University, 3-3165 Higashi-Tamagawagakuen, Machida, Tokyo 194-8543, Japan.
Organizational Affiliation: