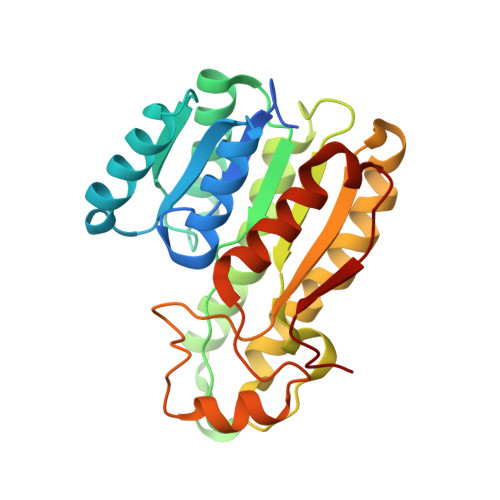
Structure-Guided Generation of a Redox-Independent Blue Fluorescent Protein from mBFP.
Seo, P.W., Jo, E.S., You, S.H., Cheong, D.E., Kim, G.J., Kim, J.S.(2019) J Mol Biology 431: 3191-3202
- PubMed: 31202883 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2019.06.005
- Primary Citation Related Structures:
6J7H, 6J7U - PubMed Abstract:
Fluorescent proteins, such as the green fluorescent protein, are used for detection of cellular components and events. However, green fluorescent protein and its derivatives have limited usage under anaerobic conditions and require a long maturation time. On the other hand, the NADPH-dependent blue fluorescent protein (BFP) without oxidative modification of residues is instantly functional in both aerobic and anaerobic systems. BFP proteins belong to a short-chain dehydrogenase/reductase (SDR) protein family, and their fluorescent property changes with reaction time in the presence of a substrate. With the aim of developing a better fluorescent reporter independent of redox state, we elucidated the crystal structure of a tetrameric mBFP from soil metagenomes with and without NADPH. Apart from the previously known regions, structure-guided mutational studies have identified several residues that contribute to the fluorescence of mBFP, including two aromatic residues (F97 and Y157) near the nicotinamide moiety of the bound NADPH. A single histidine mutation at Y157 (Y157H) has conferred more stabilized, time-independent fluorescence even in the presence of substrates. Furthermore, we discovered another SDR protein that can also emit blue fluorescence. These results open a new possibility for the development of BFP as a stable cellular reporter for widespread use, independent of subcellular environments.
- Department of Chemistry, Chonnam National University, Gwangju 61186, Republic of Korea.
Organizational Affiliation: