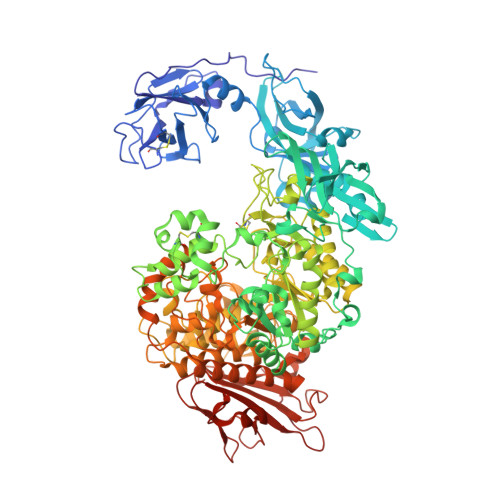
Relationship between the induced-fit loop and the activity of Klebsiella pneumoniae pullulanase.
Saka, N., Malle, D., Iwamoto, H., Takahashi, N., Mizutani, K., Mikami, B.(2019) Acta Crystallogr D Struct Biol 75: 792-803
- PubMed: 31478902 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798319010660
- Primary Citation Related Structures:
6J33, 6J34, 6J35, 6J4H - PubMed Abstract:
Klebsiella pneumoniae pullulanase (KPP) belongs to glycoside hydrolase family 13 subfamily 13 (GH13_13) and is the only enzyme that is reported to perform an induced-fit motion of the active-site loop (residues 706-710). Comparison of pullulanase structures indicated that only KPP has Leu680 present behind the loop, in contrast to the glycine found in other GH13_13 members. Analysis of the structure and activity of recombinant pullulanase from K. pneumoniae ATCC 9621 (rKPP) and its mutant (rKPP-G680L) indicated that the side chain of residue 680 is important for the induced-fit motion of the loop 706-710 and alters the binding affinity of the substrate.
- Laboratory of Applied Structural Biology, Division of Applied Life Sciences, Graduate School of Agriculture, Kyoto University, Gokasho, Uji, Kyoto 611-0011, Japan.
Organizational Affiliation: