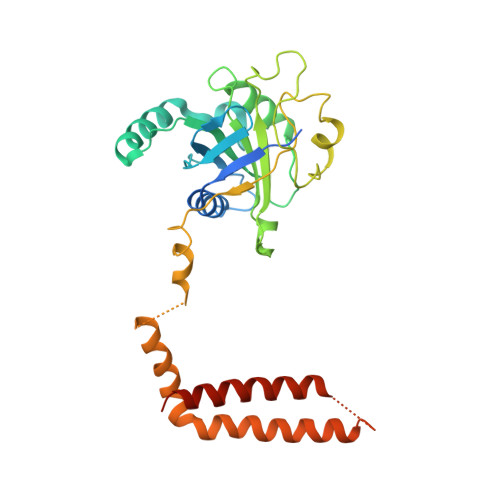
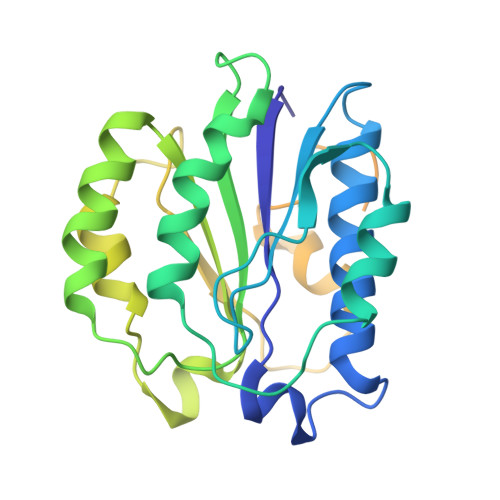
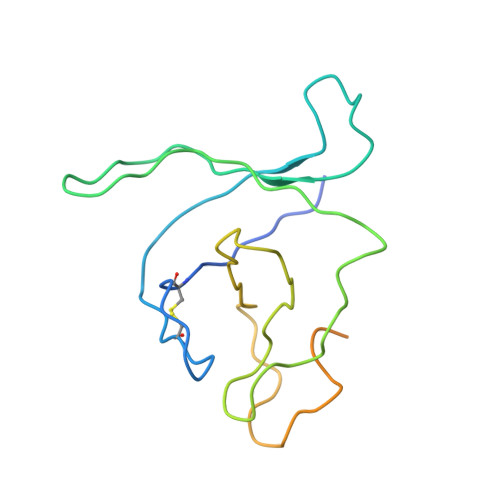
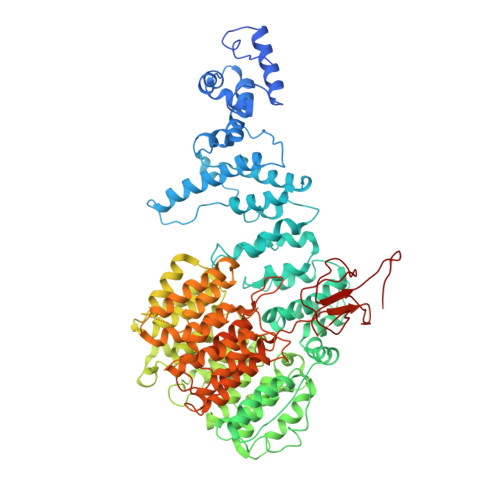
Structural Snapshots of 26S Proteasome Reveal Tetraubiquitin-Induced Conformations.
Ding, Z., Xu, C., Sahu, I., Wang, Y., Fu, Z., Huang, M., Wong, C.C.L., Glickman, M.H., Cong, Y.(2019) Mol Cell 73: 1150
- PubMed: 30792173 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2019.01.018
- Primary Citation Related Structures:
6J2C, 6J2N, 6J2Q, 6J2X, 6J30 - PubMed Abstract:
The 26S proteasome is the ATP-dependent protease responsible for regulating the proteome of eukaryotic cells through degradation of mainly ubiquitin-tagged substrates. In order to understand how proteasome responds to ubiquitin signal, we resolved an ensemble of cryo-EM structures of proteasome in the presence of K48-Ub 4 , with three of them resolved at near-atomic resolution. We identified a conformation with stabilized ubiquitin receptors and a previously unreported orientation of the lid, assigned as a Ub-accepted state C1-b. We determined another structure C3-b with localized K48-Ub 4 to the toroid region of Rpn1, assigned as a substrate-processing state. Our structures indicate that tetraUb induced conformational changes in proteasome could initiate substrate degradation. We also propose a CP gate-opening mechanism involving the propagation of the motion of the lid to the gate through the Rpn6-α2 interaction. Our results enabled us to put forward a model of a functional cycle for proteasomes induced by tetraUb and nucleotide.
- National Center for Protein Science Shanghai, State Key Laboratory of Molecular Biology, CAS Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, University of Chinese Academy of Sciences, Shanghai 201210, China; Shanghai Science Research Center, Chinese Academy of Sciences, Shanghai 201210, China.
Organizational Affiliation: