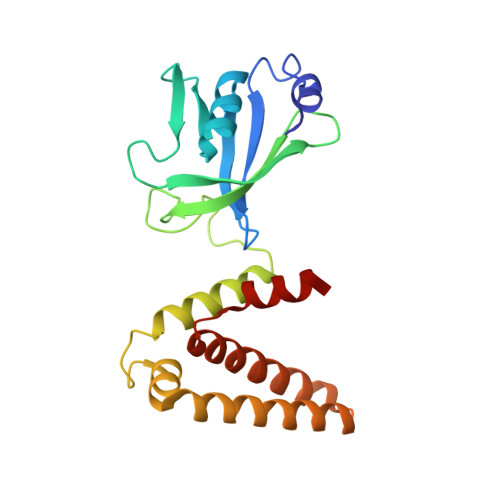
Structural analysis of Shigella flexneri bi-functional enzyme HisIE in histidine biosynthesis.
Wang, Y., Zhang, F., Nie, Y., Shang, G., Zhang, H.(2019) Biochem Biophys Res Commun 516: 540-545
- PubMed: 31235255 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.06.099
- Primary Citation Related Structures:
6J22, 6J2L - PubMed Abstract:
Histidine biosynthesis, which is absent in animals, was shown to be highly conserved among gram-negative bacteria, thus making it an attractive target for antibiotic design. There are many fusion forms of enzymes in the histidine biosynthetic pathway and people still have limited knowledge about their domain organizations and catalytic mechanisms, due to the lack of structural information. Here we report the first crystal structure of Shigella flexneri bi-functional enzyme HisIE (SfHisIE) that functions in the 2nd and 3rd steps in the histidine biosynthetic pathway. This structure shows that HisIE exists as dimers with two loops (fusion loop) connecting the individual dimer of HisE and HisI in its N-terminus and C-terminus respectively. Our mutagenesis study shows mutations in this fusion loop are lethal for bacteria indicating the advantage of gene fusion in Histidine biosynthesis. Structural analysis revealed several highly conserved residues in the putative ligand binding grooves of HisE and HisI, showing an evolutionarily conserved catalytic mechanism shared among gram negative-bacteria.
- Shanghai Institute for Advanced Immunochemical Studies (SIAIS), ShanghaiTech University, Shanghai, China.
Organizational Affiliation: