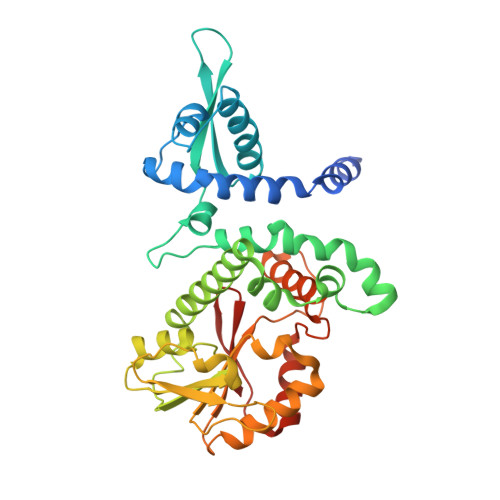
The Molecular and Structural Basis ofO-methylation Reaction in Coumarin Biosynthesis inPeucedanum praeruptorumDunn.
Zhao, Y., Wang, N., Sui, Z., Huang, C., Zeng, Z., Kong, L.(2019) Int J Mol Sci 20
- PubMed: 30934718 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms20071533
- Primary Citation Related Structures:
6IWT - PubMed Abstract:
Methoxylated coumarins represent a large proportion of officinal value coumarins while only one enzyme specific to bergaptol O -methylation (BMT) has been identified to date. The multiple types of methoxylated coumarins indicate that at least one unknown enzyme participates in the O -methylation of other hydroxylated coumarins and remains to be identified. Combined transcriptome and metabonomics analysis revealed that an enzyme similar to caffeic acid O -methyltransferase (COMT-S, S is short for similar) was involved in catalyzing all the hydroxylated coumarins in Peucedanum praeruptorum . However, the precise molecular mechanism of its substrate heterozygosis remains unsolved. Pursuing this question, we determined the crystal structure of COMT-S to clarify its substrate preference. The result revealed that Asn132, Asp271, and Asn325 govern the substrate heterozygosis of COMT-S. A single mutation, such as N132A, determines the catalytic selectivity of hydroxyl groups in esculetin and also causes production differences in bergapten. Evolution-based analysis indicated that BMT was only recently derived as a paralogue of caffeic acid O -methyltransferase (COMT) via gene duplication, occurring before the Apiaceae family divergence between 37 and 100 mya. The present study identified the previously unknown O -methylation steps in coumarin biosynthesis. The crystallographic and mutational studies provided a deeper understanding of the substrate preference, which can be used for producing specific O -methylation coumarins. Moreover, the evolutionary relationship between BMT and COMT-S was clarified to facilitate understanding of evolutionary events in the Apiaceae family.
- Key Laboratory of Bioactive Natural Product Research and State Key Laboratory of Natural Medicines, School of Traditional Chinese Pharmacy, China Pharmaceutical University, Nanjing 210009, China. zhaoyucheng1986@126.com.
Organizational Affiliation: