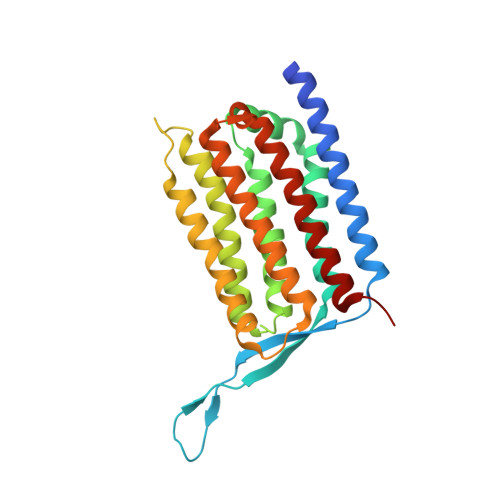
Crystal structure of heliorhodopsin.
Shihoya, W., Inoue, K., Singh, M., Konno, M., Hososhima, S., Yamashita, K., Ikeda, K., Higuchi, A., Izume, T., Okazaki, S., Hashimoto, M., Mizutori, R., Tomida, S., Yamauchi, Y., Abe-Yoshizumi, R., Katayama, K., Tsunoda, S.P., Shibata, M., Furutani, Y., Pushkarev, A., Beja, O., Uchihashi, T., Kandori, H., Nureki, O.(2019) Nature 574: 132-136
- PubMed: 31554965 Search on PubMed
- DOI: https://doi.org/10.1038/s41586-019-1604-6
- Primary Citation Related Structures:
6IS6 - PubMed Abstract:
Heliorhodopsins (HeRs) are a family of rhodopsins that was recently discovered using functional metagenomics 1 . They are widely present in bacteria, archaea, algae and algal viruses 2,3 . Although HeRs have seven predicted transmembrane helices and an all-trans retinal chromophore as in the type-1 (microbial) rhodopsin, they display less than 15% sequence identity with type-1 and type-2 (animal) rhodopsins. HeRs also exhibit the reverse orientation in the membrane compared with the other rhodopsins. Owing to the lack of structural information, little is known about the overall fold and the photoactivation mechanism of HeRs. Here we present the 2.4-Å-resolution structure of HeR from an uncultured Thermoplasmatales archaeon SG8-52-1 (GenBank sequence ID LSSD01000000). Structural and biophysical analyses reveal the similarities and differences between HeRs and type-1 microbial rhodopsins. The overall fold of HeR is similar to that of bacteriorhodopsin. A linear hydrophobic pocket in HeR accommodates a retinal configuration and isomerization as in the type-1 rhodopsin, although most of the residues constituting the pocket are divergent. Hydrophobic residues fill the space in the extracellular half of HeR, preventing the permeation of protons and ions. The structure reveals an unexpected lateral fenestration above the β-ionone ring of the retinal chromophore, which has a critical role in capturing retinal from environment sources. Our study increases the understanding of the functions of HeRs, and the structural similarity and diversity among the microbial rhodopsins.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: