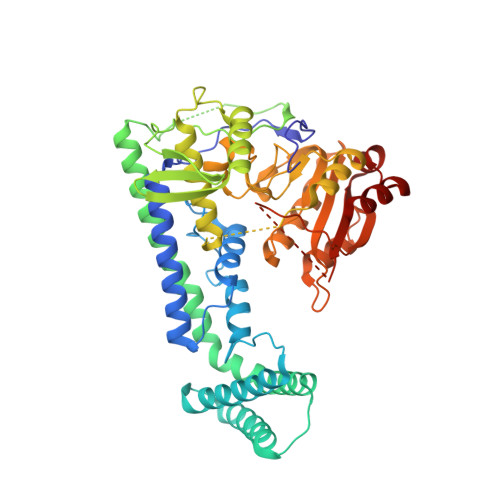
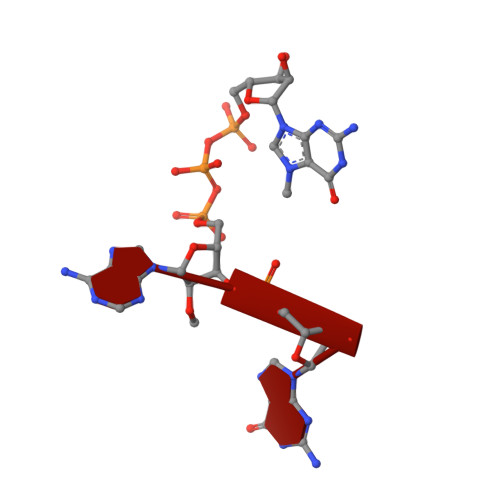
Cap-specific terminal N 6 -methylation of RNA by an RNA polymerase II-associated methyltransferase.
Akichika, S., Hirano, S., Shichino, Y., Suzuki, T., Nishimasu, H., Ishitani, R., Sugita, A., Hirose, Y., Iwasaki, S., Nureki, O., Suzuki, T.(2019) Science 363
- PubMed: 30467178 Search on PubMed
- DOI: https://doi.org/10.1126/science.aav0080
- Primary Citation Related Structures:
6IRV, 6IRW, 6IRX, 6IRY, 6IRZ, 6IS0 - PubMed Abstract:
N 6 -methyladenosine (m 6 A), a major modification of messenger RNAs (mRNAs), plays critical roles in RNA metabolism and function. In addition to the internal m 6 A, N 6 , 2'- O -dimethyladenosine (m 6 Am) is present at the transcription start nucleotide of capped mRNAs in vertebrates. However, its biogenesis and functional role remain elusive. Using a reverse genetics approach, we identified PCIF1, a factor that interacts with the serine-5-phosphorylated carboxyl-terminal domain of RNA polymerase II, as a cap-specific adenosine methyltransferase (CAPAM) responsible for N 6 -methylation of m 6 Am. The crystal structure of CAPAM in complex with substrates revealed the molecular basis of cap-specific m 6 A formation. A transcriptome-wide analysis revealed that N 6 -methylation of m 6 Am promotes the translation of capped mRNAs. Thus, a cap-specific m 6 A writer promotes translation of mRNAs starting from m 6 Am.
- Department of Chemistry and Biotechnology, Graduate School of Engineering, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-8656, Japan.
Organizational Affiliation: