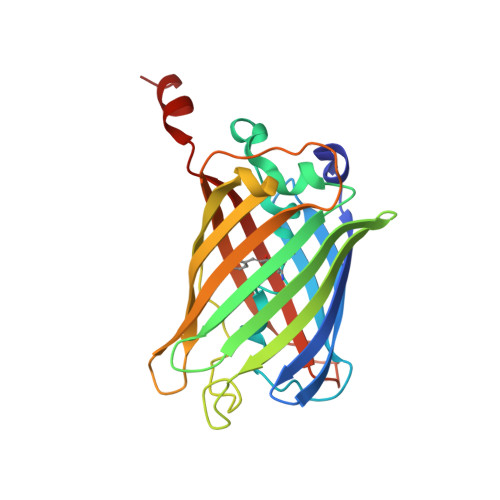
Specific modification at the C-terminal lysine residue of the green fluorescent protein variant, GFPuv, expressed in Escherichia coli.
Nakatani, T., Yasui, N., Tamura, I., Yamashita, A.(2019) Sci Rep 9: 4722-4722
- PubMed: 30886277 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-41309-8
- Primary Citation Related Structures:
6IR6, 6IR7 - PubMed Abstract:
Green fluorescent protein (GFP) is amenable to recombinant expression in various kinds of cells and is widely used in life science research. We found that the recombinant expression of GFPuv, a commonly-used mutant of GFP, in E. coli produced two distinct molecular species as judged by in-gel fluorescence SDS-PAGE. These molecular species, namely form I and II, could be separately purified by anion-exchange chromatography without any remarkable differences in the fluorescence spectra. Mass spectrometric analyses revealed that the molecular mass of form I is almost the same as the calculated value, while that of form II is approximately 1 Da larger than that of form I. Further mass spectrometric top-down sequencing pinpointed the modification in GFPuv form II, where the ε-amino group of the C-terminal Lys238 residue is converted into the hydroxyl group. No equivalent modification was observed in the native GFP in jellyfish Aequorea victoria, suggesting that this modification is not physiologically relevant. Crystal structure analysis of the two species verified the structural identity of the backbone and the vicinity of the chromophore. The modification found in this study may also be generated in other GFP variants as well as in other recombinant expression systems.
- Graduate School of Medicine, Dentistry and Pharmaceutical Sciences, Okayama University, 1-1-1, Tsushima-naka, Kita-ku, Okayama, 700-8530, Japan.
Organizational Affiliation: