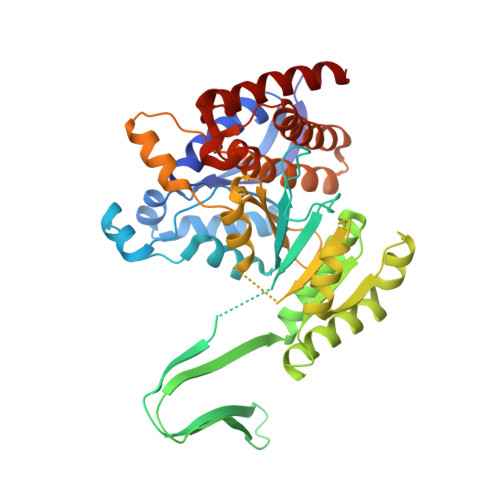
A Potent Blood-Brain Barrier-Permeable Mutant IDH1 Inhibitor Suppresses the Growth of Glioblastoma with IDH1 Mutation in a Patient-Derived Orthotopic Xenograft Model.
Machida, Y., Nakagawa, M., Matsunaga, H., Yamaguchi, M., Ogawara, Y., Shima, Y., Yamagata, K., Katsumoto, T., Hattori, A., Itoh, M., Seki, T., Nishiya, Y., Nakamura, K., Suzuki, K., Imaoka, T., Baba, D., Suzuki, M., Sampetrean, O., Saya, H., Ichimura, K., Kitabayashi, I.(2020) Mol Cancer Ther 19: 375-383
- PubMed: 31727689 Search on PubMed
- DOI: https://doi.org/10.1158/1535-7163.MCT-18-1349
- Primary Citation Related Structures:
6IO0 - PubMed Abstract:
Gliomas are the second most common primary brain tumors in adults. They are treated with combination therapies, including surgery, radiotherapy, and chemotherapy. There are currently limited treatment options for recurrent gliomas, and new targeted therapies need to be identified, especially in glioblastomas, which have poor prognosis. Isocitrate dehydrogenase (IDH) mutations are detected in various tumors, including gliomas. Most patients with IDH mutant glioma harbor the IDH1R132H subtype. Mutant IDH catalyzes the conversion of α-ketoglutarate to the oncometabolite 2-hydroxyglutarate (2-HG), which induces aberrant epigenetic status and contributes to malignant progression, and is therefore a potential therapeutic target for IDH mutant tumors. The present study describes a novel, orally bioavailable selective mutant IDH1 inhibitor, DS-1001b. The drug has high blood-brain barrier (BBB) permeability and inhibits IDH1R132H. Continuous administration of DS-1001b impaired tumor growth and decreased 2-HG levels in subcutaneous and intracranial xenograft models derived from a patient with glioblastoma with IDH1 mutation. Moreover, the expression of glial fibrillary acidic protein was strongly induced by DS-1001b, suggesting that inhibition of mutant IDH1 promotes glial differentiation. These results reveal the efficacy of BBB-permeable DS-1001b in orthotopic patient-derived xenograft models and provide a preclinical rationale for the clinical testing of DS-1001b in recurrent gliomas.
- Division of Hematological Malignancy, National Cancer Center Research Institute, Tokyo, Japan.
Organizational Affiliation: