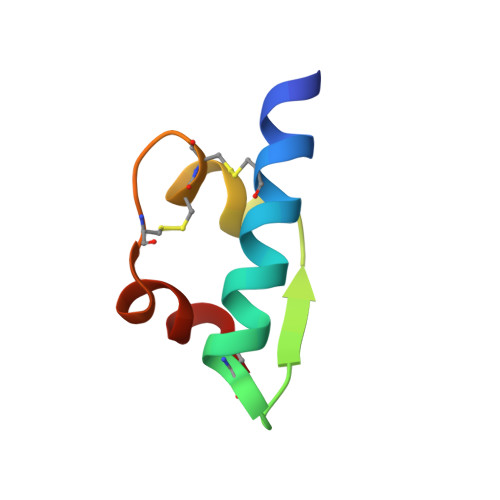
X-ray analysis of the single chain B29-A1 peptide-linked insulin molecule. A completely inactive analogue.
Derewenda, U., Derewenda, Z., Dodson, E.J., Dodson, G.G., Bing, X., Markussen, J.(1991) J Mol Biology 220: 425-433
- PubMed: 1856866 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(91)90022-x
- Primary Citation Related Structures:
6INS - PubMed Abstract:
A crystal structure of a totally inactive insulin molecule has been determined. For this insulin molecule, the first without detectable activity to be characterized, the A and B-chains are linked by a peptide bond between A1 Gly and B29 Lys. The molecule has retained all its normal self-association properties and it can also accommodate the two different conformations designated T and R, as seen in 4Zn native pig insulin crystals. The hexamers of the crosslinked insulin molecule were crystallized using the 4Zn insulin recipe of Schlichtkrull. The structure has been crystallographically refined with data extending to 2 A using restrained least-square methods. Comparison of the B29-A1 peptide crosslink insulin and the 4Zn native insulin reveals close structural similarities with the native dimer. The analysis of the structure confirms the earlier hypothesis that insulin structures in crystals are not in an active conformation and that a separation of N-terminal A-chain and C-terminal B-chain is required for interaction with the insulin receptor.
- Department of Chemistry, University of York, Heslington, U.K.
Organizational Affiliation: