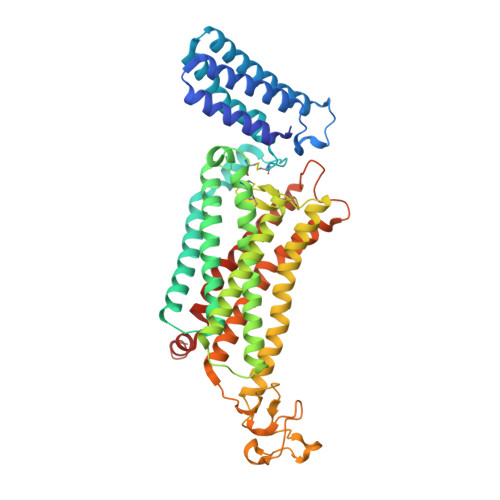
Structural basis for ligand recognition of the human thromboxane A2receptor.
Fan, H., Chen, S., Yuan, X., Han, S., Zhang, H., Xia, W., Xu, Y., Zhao, Q., Wu, B.(2019) Nat Chem Biol 15: 27-33
- PubMed: 30510189 Search on PubMed
- DOI: https://doi.org/10.1038/s41589-018-0170-9
- Primary Citation Related Structures:
6IIU, 6IIV - PubMed Abstract:
Stimulated by thromboxane A 2 , an endogenous arachidonic acid metabolite, the thromboxane A 2 receptor (TP) plays a pivotal role in cardiovascular homeostasis, and thus is considered as an important drug target for cardiovascular disease. Here, we report crystal structures of the human TP bound to two nonprostanoid antagonists, ramatroban and daltroban, at 2.5 Å and 3.0 Å resolution, respectively. The TP structures reveal a ligand-binding pocket capped by two layers of extracellular loops that are stabilized by two disulfide bonds, limiting ligand access from the extracellular milieu. These structures provide details of interactions between the receptor and antagonists, which help to integrate previous mutagenesis and SAR data. Molecular docking of prostanoid-like ligands, combined with mutagenesis, ligand-binding and functional assays, suggests a prostanoid binding mode that may also be adopted by other prostanoid receptors. These insights into TP deepen our understanding about ligand recognition and selectivity mechanisms of this physiologically important receptor.
- CAS Key Laboratory of Receptor Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: