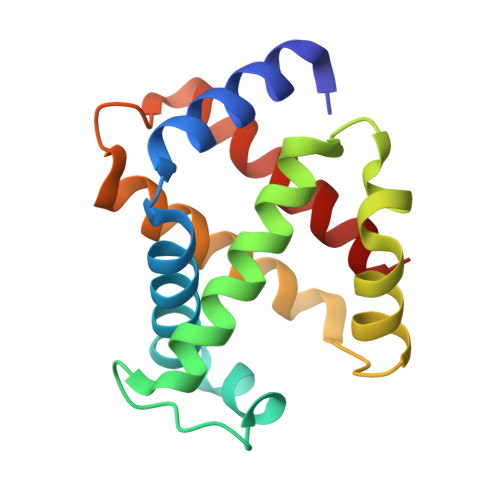
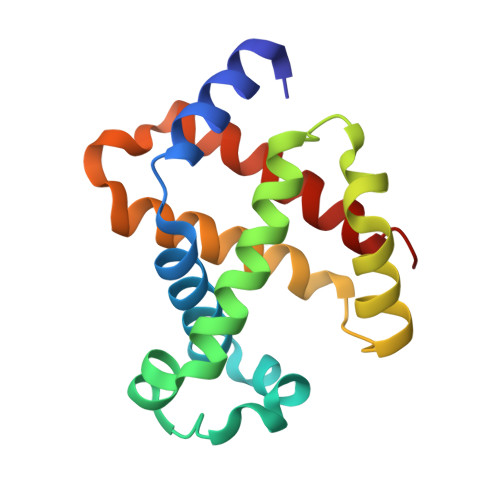
Quaternary Structure Analysis of a Hemoglobin Core in Hemoglobin-Albumin Cluster.
Morita, Y., Yamada, T., Kureishi, M., Kihira, K., Komatsu, T.(2018) J Phys Chem B 122: 12031-12039
- PubMed: 30444368 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jpcb.8b10077
- Primary Citation Related Structures:
6IHX, 6II1 - PubMed Abstract:
A core-shell ensemble of bovine hemoglobin (Hb) and human serum albumin (HSA) is an artificial O 2 carrier as a red blood cell substitute. This protein particle is created by covalent wrapping of a carbonyl Hb with HSAs: Hb R -HSA 3 cluster, where Hb R signifies the use of carbonyl Hb (relaxed (R) state conformation) as a starting material. The Hb R -HSA 3 cluster exhibits high O 2 affinity and low cooperativity. Analysis of the quaternary structure of the central Hb R in the cluster revealed that its high O 2 affinity is attributed to the physically immobile Hb R nucleus. Circular dichroism and UV-vis absorption spectroscopy showed that the structure of deoxy Hb R core closely resembles the R-state. The crystal structure of Lys-modified carbonyl Hb R was superimposed on that of carbonyl Hb. These results imply that chemical modifications of the surface Lys groups and Cys-93(β) of the carbonyl Hb with cross-linking agent interfered in the quaternary structure movement from the R-state to tense (T) state. As expected, coupling of deoxy Hb (T-state) with HSAs yielded Hb T -HSA 3 cluster having low O 2 affinity. The mixing of Hb R -HSA 3 and Hb T -HSA 3 clusters conferred a tailor-made formulation of artificial O 2 carrier with a desired O 2 affinity ( P 50 ).
- Department of Applied Chemistry, Faculty of Science and Engineering , Chuo University , 1-13-27 Kasuga , Bunkyo-ku , Tokyo 112-8551 , Japan.
Organizational Affiliation: