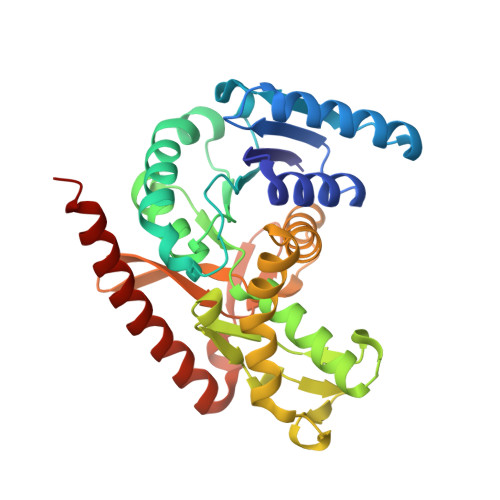
Crystal structure and biochemical characterization of malate dehydrogenase from Metallosphaera sedula
Lee, D., Hong, J., Kim, K.J.(2019) Biochem Biophys Res Commun 509: 833-838
- PubMed: 30638660 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.01.018
- Primary Citation Related Structures:
6IHD, 6IHE - PubMed Abstract:
Metallosphaera sedula is a thermoacidophilic autotrophic archaeon and known to utilize the 3-hydroxypropionate/4-hydroxybutyrate cycle (3-HP/4-HB cycle) as a carbon fixation pathway. The 3-HP/4-HB cycle in M. sedula is associated with central metabolism, and malate dehydrogenase (MDH) is an enzyme involved in the central metabolism that converts malate to oxaloacetate. To elucidate the enzymatic properties of MDH from M. sedula (MsMDH), we determined the crystal structure of MsMDH as a complex with NAD + and a ternary complex with malate and NAD + . Based on its complex structures and biochemical experiments, we observed that MsMDH can utilize both NAD + and NADP + as a cofactor. In addition, we revealed that MsMDH shows a conformational change at the active site upon substrate binding. Based on the comparison with other MDHs, we revealed that MsMDH was distinguished from general MDHs due to a Lys80 residue, and this difference is likely to influence the unique cofactor specificity of MsMDH.
- School of Life Sciences, BK21 Plus KNU Creative BioResearch Group, Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu, 702-701, Republic of Korea; KNU Institute for Microorganisms, Kyungpook National University, Daegu, 41566, Republic of Korea.
Organizational Affiliation: