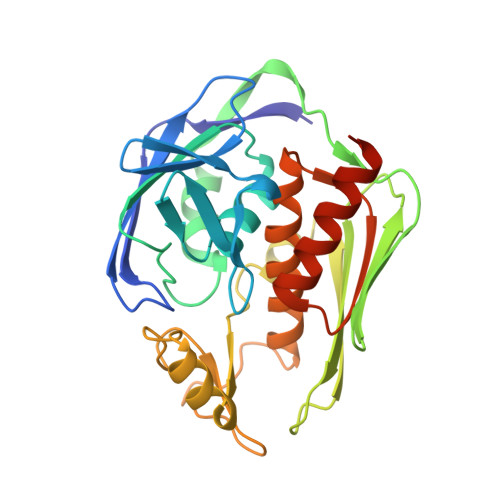
The Complex Structure of Protein AaLpxC from Aquifex aeolicus with ACHN-975 Molecule Suggests an Inhibitory Mechanism at Atomic-Level against Gram-Negative Bacteria
Fan, S., Li, D., Yan, M., Feng, X., Lv, G., Wu, G., Jin, Y., Wang, Y., Yang, Z.(2021) Molecules 26