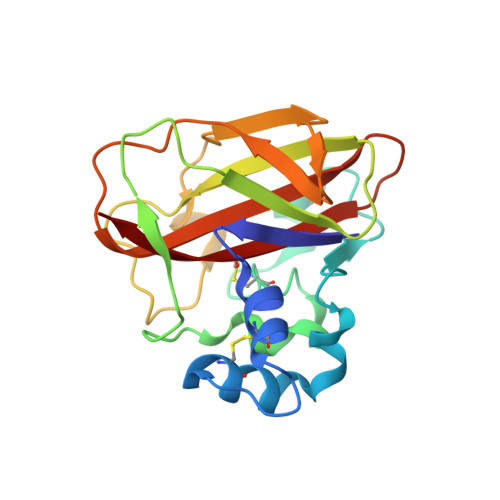
Insecticidal fern protein Tma12 is possibly a lytic polysaccharide monooxygenase.
Yadav, S.K., Singh, R., Singh, P.K., Vasudev, P.G.(2019) Planta 249: 1987-1996
- PubMed: 30903269 Search on PubMed
- DOI: https://doi.org/10.1007/s00425-019-03135-0
- Primary Citation Related Structures:
6IF7 - PubMed Abstract:
Amino acid sequence and crystal structure analyses of Tma12, an insecticidal protein isolated from the fern Tectaria macrodonta, identify it as a carbohydrate-binding protein belonging to the AA10 family of lytic polysaccharide monooxygenases, and provide the first evidence of AA10 proteins in plants. Tma12, isolated from the fern Tectaria macrodonta, is a next-generation insecticidal protein. Transgenic cotton expressing Tma12 exhibits resistance against whitefly and viral diseases. Beside its insecticidal property, the structure and function of Tma12 are unknown. This limits understanding of the insecticidal mechanism of the protein and targeted improvement in its efficacy. Here we report the amino acid sequence analysis and the crystal structure of Tma12, suggesting that it is possibly a lytic polysaccharide monooxygenase (LPMO) of the AA10 family. Amino acid sequence of Tma12 shows 45% identity with a cellulolytic LPMO of Streptomyces coelicolor. The crystal structure of Tma12, obtained at 2.2 Å resolution, possesses all the major structural characteristics of AA10 LPMOs. A H 2 O 2 -based enzymatic assay also supports this finding. It is the first report of the occurrence of LPMO-like protein in a plant. The two facts that Tma12 possesses insecticidal activity and shows structural similarity with LPMOs collectively advocate exploration of microbial LPMOs for insecticidal potential.
- Genetics and Plant Molecular Biology Division, CSIR-National Botanical Research Institute, Lucknow, 226001, India.
Organizational Affiliation: