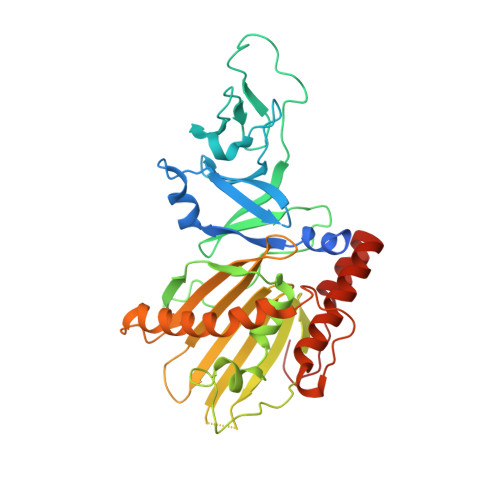
Structural and Mechanistic Insights into Caffeine Degradation by the Bacterial N-Demethylase Complex.
Kim, J.H., Kim, B.H., Brooks, S., Kang, S.Y., Summers, R.M., Song, H.K.(2019) J Mol Biology 431: 3647-3661
- PubMed: 31412262 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2019.08.004
- Primary Citation Related Structures:
6ICK, 6ICL, 6ICM, 6ICN, 6ICO, 6ICP, 6ICQ - PubMed Abstract:
Caffeine, found in many foods, beverages, and pharmaceuticals, is the most used chemical compound for mental alertness. It is originally a natural product of plants and exists widely in environmental soil. Some bacteria, such as Pseudomonas putida CBB5, utilize caffeine as a sole carbon and nitrogen source by degrading it through sequential N-demethylation catalyzed by five enzymes (NdmA, NdmB, NdmC, NdmD, and NdmE). The environmentally friendly enzymatic reaction products, methylxanthines, are high-value biochemicals that are used in the pharmaceutical and cosmetic industries. However, the structures and biochemical properties of bacterial N-demethylases remain largely unknown. Here, we report the structures of NdmA and NdmB, the initial N 1 - and N 3 -specific demethylases, respectively. Reverse-oriented substrate bindings were observed in the substrate-complexed structures, offering methyl position specificity for proper N-demethylation. For efficient sequential degradation of caffeine, these enzymes form a unique heterocomplex with 3:3 stoichiometry, which was confirmed by enzymatic assays, fluorescent labeling, and small-angle x-ray scattering. The binary structure of NdmA with the ferredoxin domain of NdmD, which is the first structural information for the plant-type ferredoxin domain in a complex state, was also determined to better understand electron transport during N-demethylation. These findings broaden our understanding of the caffeine degradation mechanism by bacterial enzymes and will enable their use for industrial applications.
- Department of Life Sciences, Korea University, 145 Anam-ro, Seongbuk-gu, Seoul 02841, Republic of Korea.
Organizational Affiliation: