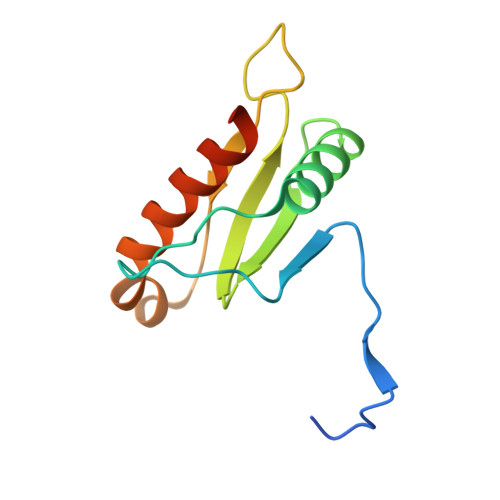
The S . Typhi effector StoD is an E3/E4 ubiquitin ligase which binds K48- and K63-linked diubiquitin.
McDowell, M.A., Byrne, A.M., Mylona, E., Johnson, R., Sagfors, A., Crepin, V.F., Lea, S., Frankel, G.(2019) Life Sci Alliance 2
- PubMed: 31142637 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.201800272
- Primary Citation Related Structures:
6IAI - PubMed Abstract:
Salmonella enterica (e.g., serovars Typhi and Typhimurium) relies on translocation of effectors via type III secretion systems (T3SS). Specialization of typhoidal serovars is thought to be mediated via pseudogenesis. Here, we show that the Salmonella Typhi STY1076/t1865 protein, named StoD, a homologue of the enteropathogenic Escherichia coli /enterohemorrhagic E. coli/Citrobacter rodentium NleG, is a T3SS effector. The StoD C terminus (StoD-C) is a U-box E3 ubiquitin ligase, capable of autoubiquitination in the presence of multiple E2s. The crystal structure of the StoD N terminus (StoD-N) at 2.5 Å resolution revealed a ubiquitin-like fold. In HeLa cells expressing StoD, ubiquitin is redistributed into puncta that colocalize with StoD. Binding assays showed that StoD-N and StoD-C bind the same exposed surface of the β-sheet of ubiquitin, suggesting that StoD could simultaneously interact with two ubiquitin molecules. Consistently, StoD interacted with both K63- (K D = 5.6 ± 1 μM) and K48-linked diubiquitin (K D = 15 ± 4 μM). Accordingly, we report the first S. Typhi-specific T3SS effector. We suggest that StoD recognizes and ubiquitinates pre-ubiquitinated targets, thus subverting intracellular signaling by functioning as an E4 enzyme.
- Sir William Dunn School of Pathology, University of Oxford, Oxford, UK.
Organizational Affiliation: