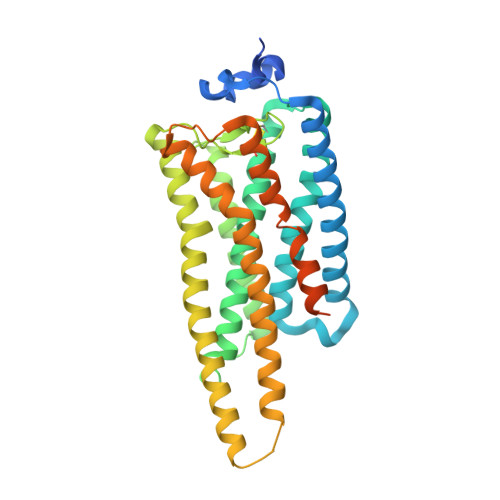
Crystal structure of jumping spider rhodopsin-1 as a light sensitive GPCR.
Varma, N., Mutt, E., Muhle, J., Panneels, V., Terakita, A., Deupi, X., Nogly, P., Schertler, G.F.X., Lesca, E.(2019) Proc Natl Acad Sci U S A 116: 14547-14556
- PubMed: 31249143 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1902192116
- Primary Citation Related Structures:
6I9K - PubMed Abstract:
Light-sensitive G protein-coupled receptors (GPCRs)-rhodopsins-absorb photons to isomerize their covalently bound retinal, triggering conformational changes that result in downstream signaling cascades. Monostable rhodopsins release retinal upon isomerization as opposed to the retinal in bistable rhodopsins that "reisomerize" upon absorption of a second photon. Understanding the mechanistic differences between these light-sensitive GPCRs has been hindered by the scarcity of recombinant models of the latter. Here, we reveal the high-resolution crystal structure of a recombinant bistable rhodopsin, jumping spider rhodopsin-1, bound to the inverse agonist 9- cis retinal. We observe a water-mediated network around the ligand hinting toward the basis of their bistable nature. In contrast to bovine rhodopsin (monostable), the transmembrane bundle of jumping spider rhodopsin-1 as well that of the bistable squid rhodopsin adopts a more "activation-ready" conformation often observed in other nonphotosensitive class A GPCRs. These similarities suggest the role of jumping spider rhodopsin-1 as a potential model system in the study of the structure-function relationship of both photosensitive and nonphotosensitive class A GPCRs.
- Department of Biology and Chemistry, Laboratory of Biomolecular Research, Paul Scherrer Institute, 5303 Villigen-PSI, Switzerland.
Organizational Affiliation: