Intrahepatic heteropolymerization of M and Z alpha-1-antitrypsin.
Laffranchi, M., Elliston, E.L., Miranda, E., Perez, J., Ronzoni, R., Jagger, A.M., Heyer-Chauhan, N., Brantly, M.L., Fra, A., Lomas, D.A., Irving, J.A.(2020) JCI Insight 5
- PubMed: 32699193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1172/jci.insight.135459
- Primary Citation Related Structures:
6I1O, 6I3Z - PubMed Abstract:
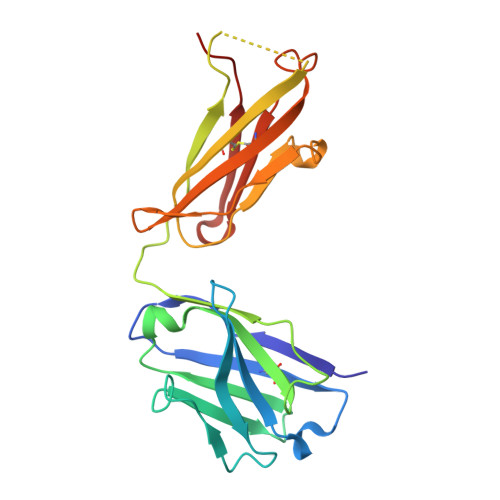
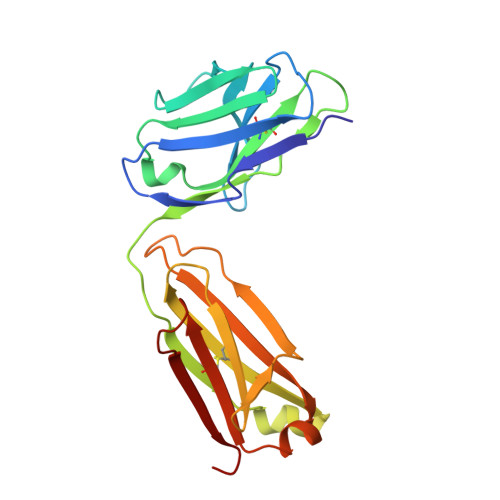
The α-1-antitrypsin (or alpha-1-antitrypsin, A1AT) Z variant is the primary cause of severe A1AT deficiency and forms polymeric chains that aggregate in the endoplasmic reticulum of hepatocytes. Around 2%-5% of Europeans are heterozygous for the Z and WT M allele, and there is evidence of increased risk of liver disease when compared with MM A1AT individuals. We have shown that Z and M A1AT can copolymerize in cell models, but there has been no direct observation of heteropolymer formation in vivo. To this end, we developed a monoclonal antibody (mAb2H2) that specifically binds to M in preference to Z A1AT, localized its epitope using crystallography to a region perturbed by the Z (Glu342Lys) substitution, and used Fab fragments to label polymers isolated from an MZ heterozygote liver explant. Glu342 is critical to the affinity of mAb2H2, since it also recognized the mild S-deficiency variant (Glu264Val) present in circulating polymers from SZ heterozygotes. Negative-stain electron microscopy of the Fab2H2-labeled liver polymers revealed that M comprises around 6% of the polymer subunits in the MZ liver sample. These data demonstrate that Z A1AT can form heteropolymers with polymerization-inert variants in vivo with implications for liver disease in heterozygous individuals.
- Department of Molecular and Translational Medicine, University of Brescia, Brescia, Italy.
Organizational Affiliation: