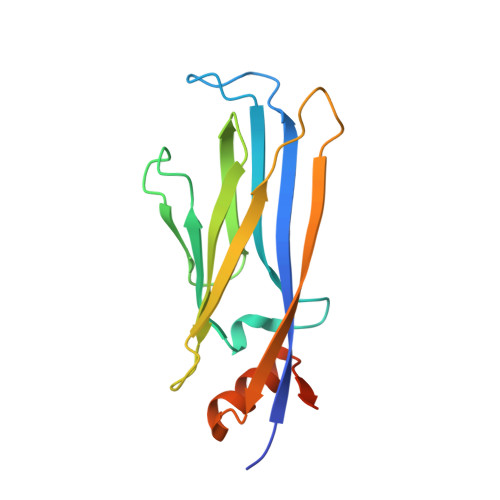
Structure-Based Approach toward Identification of Inhibitory Fragments for Eleven-Nineteen-Leukemia Protein (ENL).
Heidenreich, D., Moustakim, M., Schmidt, J., Merk, D., Brennan, P.E., Fedorov, O., Chaikuad, A., Knapp, S.(2018) J Med Chem 61: 10929-10934
- PubMed: 30407816 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01457
- Primary Citation Related Structures:
6HPW, 6HPX, 6HPY, 6HPZ, 6HQ0 - PubMed Abstract:
Lysine acetylation is an epigenetic mark that is principally recognized by bromodomains, and recently structurally diverse YEATS domains also emerged as readers of lysine acetyl/acylations. Here we present a crystallography-based strategy and the discovery of fragments binding to the ENL YEATS domain, a potential drug target. Crystal structures combined with synthetic efforts led to the identification of a submicromolar binder, providing first starting points for the development of chemical probes for this reader domain family.
- Institute of Pharmaceutical Chemistry , Goethe-University Frankfurt , 60438 Frankfurt , Germany.
Organizational Affiliation: