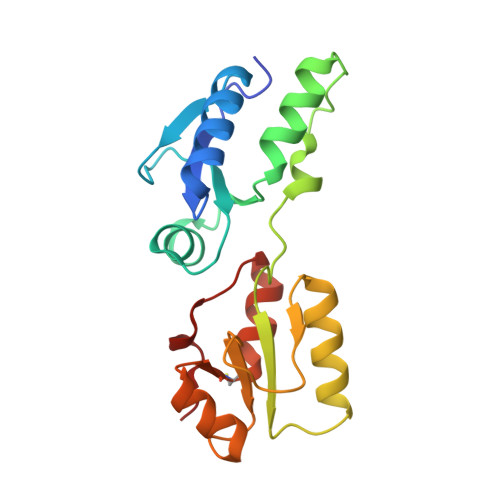
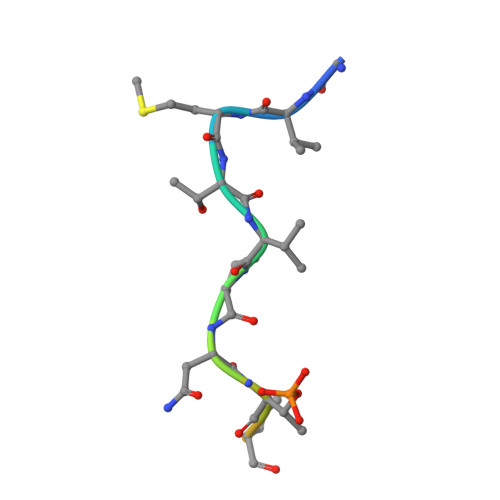
BRCT domains of the DNA damage checkpoint proteins TOPBP1/Rad4 display distinct specificities for phosphopeptide ligands.
Day, M., Rappas, M., Ptasinska, K., Boos, D., Oliver, A.W., Pearl, L.H.(2018) Elife 7
- PubMed: 30295604 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.39979
- Primary Citation Related Structures:
6HM3, 6HM4, 6HM5 - PubMed Abstract:
TOPBP1 and its fission yeast homologueRad4, are critical players in a range of DNA replication, repair and damage signalling processes. They are composed of multiple BRCT domains, some of which bind phosphorylated motifs in other proteins. They thus act as multi-point adaptors bringing proteins together into functional combinations, dependent on post-translational modifications downstream of cell cycle and DNA damage signals. We have now structurally and/or biochemically characterised a sufficient number of high-affinity complexes for the conserved N-terminal region of TOPBP1 and Rad4 with diverse phospho-ligands, including human RAD9 and Treslin, and Schizosaccharomyces pombe Crb2 and Sld3, to define the determinants of BRCT domain specificity. We use this to identify and characterise previously unknown phosphorylation-dependent TOPBP1/Rad4-binding motifs in human RHNO1 and the fission yeast homologue of MDC1, Mdb1. These results provide important insights into how multiple BRCT domains within TOPBP1/Rad4 achieve selective and combinatorial binding of their multiple partner proteins.
- Cancer Research UK DNA Repair Enzymes Group, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, United Kingdom.
Organizational Affiliation: