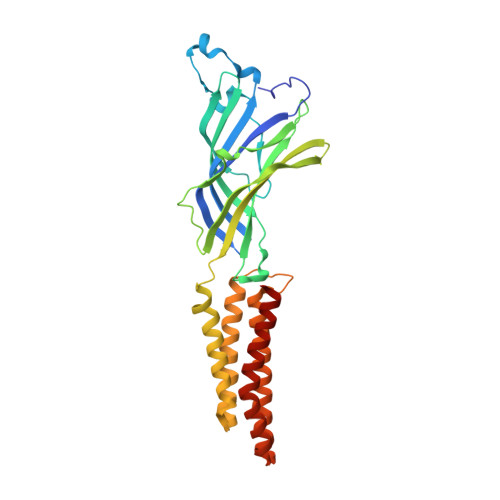
Structural evidence for the binding of monocarboxylates and dicarboxylates at pharmacologically relevant extracellular sites of a pentameric ligand-gated ion channel.
Fourati, Z., Sauguet, L., Delarue, M.(2020) Acta Crystallogr D Struct Biol 76: 668-675
- PubMed: 32627739 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S205979832000772X
- Primary Citation Related Structures:
6HJ3, 6HJA, 6HJB, 6HJI, 6HJZ, 6HPP - PubMed Abstract:
GLIC is a bacterial homologue of the pentameric ligand-gated ion channels (pLGICs) that mediate the fast chemical neurotransmission of nerve signalling in eukaryotes. Because the activation and allosteric modulation features are conserved among prokaryotic and eukaryotic pLGICs, GLIC is commonly used as a model to study the allosteric transition and structural pharmacology of pLGICs. It has previously been shown that GLIC is inhibited by some carboxylic acid derivatives. Here, experimental evidence for carboxylate binding to GLIC is provided by solving its X-ray structures with a series of monocarboxylate and dicarboxylate derivatives, and two carboxylate-binding sites are described: (i) the `intersubunit' site that partially overlaps the canonical pLGIC orthosteric site and (ii) the `intrasubunit' vestibular site, which is only occupied by a subset of the described derivatives. While the intersubunit site is widely conserved in all pLGICs, the intrasubunit site is only conserved in cationic eukaryotic pLGICs. This study sheds light on the importance of these two extracellular modulation sites as potential drug targets in pLGICs.
- Unité Dynamique Structurale des Macromolécules, Institut Pasteur, 25 Rue du Docteur Roux, F-75015 Paris, France.
Organizational Affiliation: